Advertisement
Research ArticleGastroenterologyOncology Free access | 10.1172/JCI125581
H. pylori infection alters repair of DNA double-strand breaks via SNHG17
Taotao Han,1 Xiaohui Jing,1 Jiayu Bao,1 Lianmei Zhao,1,2 Aidong Zhang,1 Renling Miao,1 Hui Guo,1 Baoguo Zhou,3 Shang Zhang,1 Jiazeng Sun,1 and Juan Shi1
1Department of Biochemistry and Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2Research Center, Fourth Affiliated Hospital of Hebei Medical University, Shijiazhuang, China.
3Department of General Surgery, First Affiliated Hospital of Harbin Medical University, Heilongjiang, China.
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
Authorship note: TH, XJ, and JB contributed equally to this work.
Find articles by Han, T. in: JCI | PubMed | Google Scholar
1Department of Biochemistry and Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2Research Center, Fourth Affiliated Hospital of Hebei Medical University, Shijiazhuang, China.
3Department of General Surgery, First Affiliated Hospital of Harbin Medical University, Heilongjiang, China.
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
Authorship note: TH, XJ, and JB contributed equally to this work.
Find articles by Jing, X. in: JCI | PubMed | Google Scholar
1Department of Biochemistry and Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2Research Center, Fourth Affiliated Hospital of Hebei Medical University, Shijiazhuang, China.
3Department of General Surgery, First Affiliated Hospital of Harbin Medical University, Heilongjiang, China.
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
Authorship note: TH, XJ, and JB contributed equally to this work.
Find articles by
Bao, J.
in:
JCI
|
PubMed
|
Google Scholar
|

1Department of Biochemistry and Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2Research Center, Fourth Affiliated Hospital of Hebei Medical University, Shijiazhuang, China.
3Department of General Surgery, First Affiliated Hospital of Harbin Medical University, Heilongjiang, China.
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
Authorship note: TH, XJ, and JB contributed equally to this work.
Find articles by Zhao, L. in: JCI | PubMed | Google Scholar
1Department of Biochemistry and Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2Research Center, Fourth Affiliated Hospital of Hebei Medical University, Shijiazhuang, China.
3Department of General Surgery, First Affiliated Hospital of Harbin Medical University, Heilongjiang, China.
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
Authorship note: TH, XJ, and JB contributed equally to this work.
Find articles by Zhang, A. in: JCI | PubMed | Google Scholar
1Department of Biochemistry and Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2Research Center, Fourth Affiliated Hospital of Hebei Medical University, Shijiazhuang, China.
3Department of General Surgery, First Affiliated Hospital of Harbin Medical University, Heilongjiang, China.
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
Authorship note: TH, XJ, and JB contributed equally to this work.
Find articles by Miao, R. in: JCI | PubMed | Google Scholar
1Department of Biochemistry and Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2Research Center, Fourth Affiliated Hospital of Hebei Medical University, Shijiazhuang, China.
3Department of General Surgery, First Affiliated Hospital of Harbin Medical University, Heilongjiang, China.
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
Authorship note: TH, XJ, and JB contributed equally to this work.
Find articles by Guo, H. in: JCI | PubMed | Google Scholar
1Department of Biochemistry and Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2Research Center, Fourth Affiliated Hospital of Hebei Medical University, Shijiazhuang, China.
3Department of General Surgery, First Affiliated Hospital of Harbin Medical University, Heilongjiang, China.
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
Authorship note: TH, XJ, and JB contributed equally to this work.
Find articles by Zhou, B. in: JCI | PubMed | Google Scholar
1Department of Biochemistry and Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2Research Center, Fourth Affiliated Hospital of Hebei Medical University, Shijiazhuang, China.
3Department of General Surgery, First Affiliated Hospital of Harbin Medical University, Heilongjiang, China.
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
Authorship note: TH, XJ, and JB contributed equally to this work.
Find articles by Zhang, S. in: JCI | PubMed | Google Scholar
1Department of Biochemistry and Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2Research Center, Fourth Affiliated Hospital of Hebei Medical University, Shijiazhuang, China.
3Department of General Surgery, First Affiliated Hospital of Harbin Medical University, Heilongjiang, China.
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
Authorship note: TH, XJ, and JB contributed equally to this work.
Find articles by Sun, J. in: JCI | PubMed | Google Scholar
1Department of Biochemistry and Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China.
2Research Center, Fourth Affiliated Hospital of Hebei Medical University, Shijiazhuang, China.
3Department of General Surgery, First Affiliated Hospital of Harbin Medical University, Heilongjiang, China.
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
Authorship note: TH, XJ, and JB contributed equally to this work.
Find articles by Shi, J. in: JCI | PubMed | Google Scholar
Authorship note: TH, XJ, and JB contributed equally to this work.
Published June 15, 2020 - More info
J Clin Invest. 2020;130(7):3901–3918. https://doi.org/10.1172/JCI125581.
© 2020 American Society for Clinical Investigation
Received: November 15, 2018; Accepted: April 14, 2020
-
Abstract
Chronic infections can lead to carcinogenesis through inflammation-related mechanisms. Chronic infection of the human gastric mucosa with Helicobacter pylori is a well-known risk factor for gastric cancer. However, the mechanisms underlying H. pylori–induced gastric carcinogenesis are incompletely defined. We aimed to screen and clarify the functions of long noncoding RNAs (lncRNAs) that are differentially expressed in H. pylori–related gastric cancer. We found that lncRNA SNHG17 was upregulated by H. pylori infection and markedly increased the levels of double-strand breaks (DSBs). SNHG17 overexpression correlated with poor overall survival in patients with gastric cancer. The recruitment of NONO by overabundant nuclear SNHG17, along with the role of cytoplasmic SNHG17 as a decoy for miR-3909, which regulates Rad51 expression, shifted the DSB repair balance from homologous recombination toward nonhomologous end joining. Notably, during chronic H. pylori infection, SNHG17 knockdown inhibited chromosomal aberrations. Our findings suggest that spatially independent deregulation of the SNHG17/NONO and SNHG17/miR-3909/RING1/Rad51 pathways upon H. pylori infection promotes tumorigenesis in gastric cancer by altering the DNA repair system, which is critical for the maintenance of genomic stability. Upregulation of SNHG17 by H. pylori infection might be an undefined link between cancer and inflammation.
-
Introduction
Gastric cancer is among the most common malignancies; it is the third leading cause of cancer-associated mortality worldwide and is a serious threat to public health (1). Many factors are implicated in the carcinogenesis and progression of gastric cancer, including environmental risk factors, genetic alterations, and host factors. Helicobacter pylori is a well-known risk factor for gastric cancer (1). In 1994, the International Agency for Research on Cancer (IARC) defined H. pylori as a class I carcinogen strongly associated with gastric cancer (2). Chronic and persistent inflammation contributes to the initiation and progression of some kinds of tumors, and H. pylori infection can stimulate chronic inflammation in the gastric mucosa, which induces the normal gastric epithelium to progress through a series of well-defined steps into carcinoma (3). Epidemiological studies have confirmed the relationship between H. pylori infection and gastric cancer, and this relationship has also been demonstrated experimentally in rodent models using C57BL/6 mice and Mongolian gerbils (3–5). However, the mechanism by which H. pylori infection mediates gastric cancer progression is incompletely defined.
Gastric cancer develops after H. pylori infection via the long-term accumulation of molecular alterations. H. pylori infection induces chronic inflammation to trigger genetic alterations along with DNA damage in gastric epithelial cells (6, 7). Recent global analyses of gastric cancer genomes have provided valuable insight into the molecular mechanisms underlying gastric carcinogenesis. A study from The Cancer Genome Atlas (TCGA) Research Network based on 295 primary gastric adenocarcinomas defined 4 major genomic subtypes of gastric cancer: EBV-infected tumors (9%), microsatellite instability (MSI) tumors (22%), genomically stable tumors (20%), and chromosomally unstable tumors (50%). These classifications suggest that genomic instability is important in gastric cancer progression (8).
Many pathogenic bacteria have been shown to cause host cell DNA damage, often resulting in DNA double-strand breaks (DSBs) (9, 10). Abnormalities in DNA damage repair may result in genomic instability (11, 12). To manage these DSB-induced lesions, eukaryotic cells have developed an intricate and efficient DNA repair system. Homologous recombination (HR) and nonhomologous end joining (NHEJ) are the major molecular pathways responsible for DSB repair (13, 14). HR is an error-free repair pathway that uses homologous DNA sequences as templates. In contrast, NHEJ is an error-prone pathway initiated by recruitment of the DNA-PKcs complex to the DNA break ends. NHEJ joins 2 DNA ends that are independent of the chromosome locus. This type of repair occasionally results in chromosomal abnormalities, such as chromosomal deletions and translocations. Therefore, NHEJ repair is relatively error prone and responsible for genome rearrangement (14). Thus, efficient and intricate repair of DSBs is crucial for the maintenance of genomic integrity.
Many studies have reported that H. pylori infection induces DNA damage and regulates DNA repair (15–19). These findings were further supported by similar results in H. pylori–infected patients (20, 21). The regulation of gene expression by long noncoding RNAs (lncRNAs) at the epigenetic, transcriptional, and posttranscriptional levels has been widely studied (22). lncRNAs are involved in many important pathological processes, such as cell differentiation, cell death, and individual development (22). Many lncRNAs that function as oncogenes or tumor suppressors have been reported recently, suggesting that lncRNAs may be useful in the future as diagnostic, prognostic, and therapeutic tumor biomarkers (23). Recently, numerous studies have shown the dysregulation of lncRNAs in gastric cancer. For example, H19 and uc001lsz play important roles in gastric cancer and might be useful as biomarkers for the diagnosis of early gastric cancer (24, 25). In addition, lncRNA HOXA11-AS was reported to enhance cell proliferation and migration in gastric cancer by scaffolding the chromatin regulators EZH2, LSD1, and DNMT1 (26). We hypothesized that many unidentified lncRNAs are dysregulated in gastric cancer progression, especially in gastric cancer related to H. pylori infection. To investigate the mechanism underlying the association between H. pylori infection and genomic instability, we screened lncRNAs that are dysregulated in H. pylori–related gastric cancer. We identified lncRNA-SNHG17, which was upregulated by H. pylori infection and functionally increased the DSB levels. The recruitment of NONO by overabundant nuclear SNHG17, along with the role of cytoplasmic SNHG17 as a decoy for miR-3909, which regulates RING1/Rad51 expression, shifted the DSB repair balance from HR to NHEJ. These results indicate that spatially independent deregulation of the SNHG17/NONO and SNHG17/miR-3909/Rad51 pathways promotes gastric cancer progression by altering the DNA repair system, which is critical for the maintenance of genomic stability.
-
Results
Identification of H. pylori infection–induced lncRNA SNHG17. To identify lncRNAs associated with H. pylori infection, we analyzed lncRNA and mRNA expression in GES-1 normal gastric epithelial cells infected with the H. pylori standard strain NCTC11637. Three samples of H. pylori–negative normal gastric epithelial tissues, 6 samples of H. pylori–positive gastritis tissues, and 4 samples of H. pylori–positive gastric cancer tissues were collected for the microarray analysis of lncRNAs and mRNAs. As shown in Figure 1, A and B, systematic variations were screened in the expression of lncRNAs related to H. pylori infection status. To validate these results, quantitative real-time PCR (qRT-PCR) analysis of 6 randomly selected lncRNAs from among the differentially expressed lncRNAs in H. pylori–infected GES-1 cells confirmed that the trend in the expression levels of these lncRNAs was consistent with the microarray results (Supplemental Figure 1A; supplemental material available online with this article; https://doi.org/10.1172/JCI125581DS1). Some statistically marked overlapping dysregulated lncRNAs in the National Center for Biotechnology Information’s (NCBI) Gene Expression Omnibus (GEO) GSE111762 and GSE111763 microarray data sets were identified (Supplemental Table 1A).
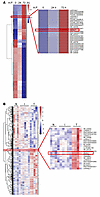 Figure 1
Figure 1Identification of the H. pylori infection–induced lncRNA SNHG17. (A) Heatmap of lncRNAs expressed differentially in the nonmalignant gastric epithelial cell line GES-1 at 24 hours and 72 hours after infection with H. pylori (H.P.) standard strain NCTC11637 (MOI, 200:1). LncRNAs with expression change fold over 2 (0 hours vs. 72 hours) and gradually upregulated or downregulated during the process of H. pylori infection are visualized here using color from low (navy blue) to high (brick red). (B) Heatmap of lncRNAs expressed differentially in H. pylori–negative normal gastric epithelial tissues, H. pylori–positive gastritis tissues, and H. pylori–positive gastric cancer tissues. lncRNAs with expression change fold over 2 (normal group vs. tumor group) and gradually upregulated or downregulated with increased gastric mucosal lesion degree were screened. N, normal group; I, inflammation group; T, tumor group.
Aberrations in DNA damage repair are an important cause of genomic instability, a characteristic of most cancers (27, 28). To identify the lncRNAs related to H. pylori infection–induced genomic instability, genomic instability–related mRNAs differentially expressed in H. pylori–infected cells were selected for clustering with deregulated lncRNAs. The degree of coexpression, which is defined as the number of directly linked neighbors of an lncRNA, was ranked (Supplemental Table 1B). As shown in Supplemental Figure 1B, several lncRNAs were located at central nodes of the network. Therefore, based on the comparison of the overlapping lncRNAs in the 2 microarrays and the coexpression network, SNHG17 was selected, as this lncRNA was not only upregulated upon H. pylori infection, but also closely associated with genomic stability.
Characterization of SNHG17 expression in gastric tissues. qRT-PCR analysis showed that the expression level of SNHG17 was gradually upregulated with increased gastric mucosal lesion degree (Figure 2A). In addition, ISH in paraffin-embedded tissue sections revealed a gradual increase in the expression level of SNHG17, with an increase in the gastric mucosal lesion degree.
 Figure 2
Figure 2SNHG17 expression was increased by H. pylori infection. (A) qRT-PCR analysis of SNHG17 expression in normal stomach mucosa tissues (NG) (n = 10), intestinal metaplasia tissues (IM) (n = 15), dysaplasia tissues (Dys) ( n = 15), and gastric cancer tissues (AdCa) (n = 112). (B) Comparison of SNHG17 expression between H. pylori–negative and H. pylori–positive tissues. (C) SNHG17 expression was analyzed by qRT-PCR in gastric cancer samples and adjacent nontumor gastric tissues (n = 112). (D) Low and high SNHG17 expression in gastric cancer tissues and adjacent nontumor gastric tissues were analyzed based on the data above. (E) SNHG17 expression detected by ISH in paraffin-embedded tissue sections. N, normal tissues; T, tumor tissues. Scale bars: 200 μm. Data are represented as mean ± SEM. *P < 0.05; **P < 0.01, ANOVA (A); 2-tailed Student’s t test (B and C).
Moreover, we performed qRT-PCR analysis on 112 tumor/matched normal gastric cancer samples. Comparison between the H. pylori–negative and H. pylori–positive tissues showed a marked enhancement of SNHG17 expression upon H. pylori infection (Figure 2B). The results also confirmed that SNHG17 expression was significantly higher in cancerous tissues (Figure 2C) than in normal tissues and was upregulated in 74.11% of the gastric cancer patients (Figure 2D). No significant correlation was found between SNHG17 expression and the age or sex of the patients or the gastric cancer tissue differentiation degree. However, the SNHG17 expression level was correlated with the tumor node metastasis (TNM) stage (P = 0.013) and H. pylori infection (P = 0.034) (Supplemental Table 2). ISH in paraffin-embedded tissues further revealed higher SNHG17 expression in gastric cancer tissues than in adjacent tissues (Figure 2E). These results suggest that SNHG17 plays an important role in the progression of gastric cancer in addition to participating in the progression of stages upon H. pylori infection.
Furthermore, the SNHG17 expression level was markedly upregulated in gastric cancer tissue relative to that in normal tissue (Supplemental Figure 1D). Given that SNHG17 is a mature splicing transcript of a small nucleolar RNA (snoRNA) SNORA host gene, these snoRNAs were also analyzed based on the TCGA database. As shown in Supplemental Figure 1D, the expression levels of these snoRNAs were significantly increased in gastric cancer. Analysis of an additional public data set (GSE84787) further validated the upregulation of SNHG17 in gastric cancer samples versus matched normal gastric tissues (Supplemental Figure 1E). Kaplan-Meier Plotter (29) (http://kmplot.com/analysis) analysis revealed that the overall survival of gastric cancer patients with high SNHG17 expression was significantly poorer than that of gastric cancer patients with low SNHG17 expression (Supplemental Figure 1F). In conclusion, SNHG17 is an H. pylori–induced lncRNA and might be required for gastric cancer progression.
SNHG17 expression was increased by H. pylori infection. To determine whether H. pylori infection alters SNHG17 expression in vitro, we first measured SNHG17 expression after coculture of H. pylori with GES-1 cells and SGC-7901 gastric cancer cells. SNHG17 expression increased with H. pylori infection in a time- and dose-dependent manner (Figure 3A). In addition, the expression of intronic snoRNAs was upregulated with H. pylori infection (Supplemental Figure 2A).
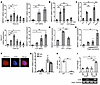 Figure 3
Figure 3SNHG17 expression was increased by H. pylori infection. (A) qRT-PCR analysis of SNHG17 expression in GES-1 (top) and SGC-7901 (bottom) infected with H. pylori for the indicated times with MOI 100:1 or for indicated MOIs with 6 hours. (B) qRT-PCR analysis of SNHG17 expression in GES-1 cells infected with “dead” H. pylori for 24 hours. Ctrl, control. (C) SGC-7901 cells were infected with H. pylori for 6 hours, after which bacteria were eradicated by antibiotic therapy and cells were allowed to recover for 24 hours. Then, qRT-PCR analysis of SNHG17 expression was performed. ET, eradication therapy. (D) RT-PCR analysis of SNHG17 expression in GES-1 cells infected with ΔcagA H. pylori for 24 hours. (E) CagA overexpression plasmid transfection into GES-1 cells induced increasing SNHG17 expression. (F) Fluorescence ISH of GES-1 cells performed using a Cy3-labeled SNHG17 probe (red). Nuclei were counterstained using DAPI (blue). Scale bars: 10 μm. Samples were assayed 3 times. (G) qRT-PCR analysis of SNHG17 expression in cytoplasmic (Cyto) and nuclear (Nuc) fractions of RNA in GES-1 cells infected with H. pylori. (H) Luciferase (LUC) activity in 293T cells transfected with the reporter construct as indicated. Mut, mutated. (I) ChIP-PCR analysis of p65-binding activity on the predicted sites in GES-1 cells infected with H. pylori. Data are represented as mean ± SEM. *P < 0.05; **P < 0.01, ANOVA (A–E and I); 2-tailed Student’s t test (H).
To investigate whether the effect of increasing SNHG17 expression is specific to viable H. pylori, we compared this capability between ethanol-killed and viable H. pylori. Only viable H. pylori efficiently induced SNHG17 expression, as determined by qRT-PCR (Figure 3B). Again, GES-1 cells were infected with H. pylori for 6 hours, bacteria were then eradicated by antibiotic therapy, and the cells were allowed to recover for 48 hours. Antibiotic eradication resulted in the recovery of SNHG17 expression to the baseline level (Figure 3C). These data revealed that the ability to increase SNHG17 expression was specific to viable H. pylori.
The pathogenicity of H. pylori is mainly due to its multiple virulence components, especially the most widely studied H. pylori virulence factor, CagA (26). We further revealed that the expression of SNHG17 was upregulated when cells were cocultured with clinically isolated strains positive for CagA expression (Supplemental Figure 2B). We next examined the ability of isogenic gene deletion mutants in the NCTC11637 strain background lacking the entire island (ΔcagPAI) to induce SNHG17 expression. As shown in Figure 3D, mutants lacking the entire cagPAI exhibited no evident ability to induce SNHG17 expression. Moreover, transfection of a CagA overexpression plasmid induced SNHG17 expression in GES-1 cells (Figure 3E), suggesting that CagA might be the critical virulence factor inducing the expression of SNHG17.
Characterization of SNHG17. Bioinformatics analysis confirmed that SNHG17 has no coding capacity (http://genome.ucsc.edu/) (Supplemental Figure 3A). According to the NCBI database, SNHG17 exists as several splice variant transcripts with a poly(A) tail structure. Primers were designed according to the NCBI sequence to amplify the full-length transcript of SNHG17. RT-PCR analysis of GES-1 cells infected with H. pylori revealed that the expression of SNHG17 resulted in 3 transcripts of different lengths (Supplemental Table 3). Among these transcripts, the full-length 818-nt transcript was the most abundant in GES-1 cells infected with H. pylori based on TA clone sequencing analysis. Therefore, the function of the 818-nt transcript was investigated in this study. The subcellular localization of SNHG17 was detected by RNA FISH. Cells treated with probe pools targeting SNHG17 exhibited staining in both the nuclear and cytoplasmic compartments (Figure 3F), which was further confirmed by RT-PCR analysis of SNHG17 in the nuclear and cytosolic fractions (Supplemental Figure 3B). We next quantified SNHG17 expression in the cytoplasmic and nuclear fractions of RNA from cells infected with H. pylori. SNHG17 expression increased in both the cytoplasmic and nuclear fractions of cells upon H. pylori infection, and the upregulation of SNHG17 among cytoplasmic RNA was more marked than that among nuclear RNA, indicating that both nuclear and cytoplasmic SNHG17 might play roles in H. pylori infection (Figure 3G).
The expression of SNHG17 was higher in a panel of gastric cancer cell lines than in GES-1 normal gastric epithelial cells (Supplemental Figure 3C). The SGC-7901 and AGS cell lines exhibited the highest expression of SNHG17 among these cell lines; therefore, in the following studies, we investigated the function of SNHG17 in SGC-7901 and AGS cells.
H. pylori activates various critical pathways, including NF-κB, β-catenin, and PI3K/AKT pathways (26–28), in gastric epithelial cells. Among these pathways, NF-κB pathway activation plays a critical role in the H. pylori–induced transformation from inflammation to cancer. In the present study, we demonstrated the activation of NF-κB upon H. pylori infection (Supplemental Figure 3D). Given that a previous ChIP-Seq assay with NF-κB p65 showed a strong signal at the SNHG17 locus (Supplemental Figure 3E), we concluded that NF-κB activation may participate in maintaining the high level of SNHG17 expression in H. pylori–infected cells. We identified an imperfect κB element spanning nt positions +509 to +518 with the sequence 5′-GGAAGCCTCC-3′ in intron 1 of the human SNHG17 gene. To confirm the activity of the binding sites, luciferase reporter plasmids containing WT and mutated NF-κB (NF-κB mut) binding sites were constructed (Supplemental Figure 3F). The reporter activity was decreased by single mutation of the NF-κB–binding site (Figure 3H), suggesting that NF-κB might promote the transcription of SNHG17. The results of the ChIP assays showed that the binding of NF-κB p65 to the SNHG17 promoter was significantly increased in H. pylori–stimulated GES-1 cells compared with that in unstimulated GES-1 cells (Figure 3I).
SNHG17 expression increased the accumulation of DSBs upon H. pylori infection and shifted DSB repair toward NHEJ. The results of the lncRNA-mRNA coexpression network analysis revealed that SNHG17 was coexpressed with many genomic stability–related mRNAs, indicating that SNHG17 might mediate H. pylori infection–induced DSBs. First, we depleted SNHG17 by siRNA in SGC-7901 cells (Supplemental Figure 4A). Knockdown of SNHG17 induced a decrease in the levels of phospho–histone H2AX (γ-H2AX), a marker of DSBs (Supplemental Figure 4B); this finding was supported by Western blot analysis results (Supplemental Figure 4C).
We further constructed SGC-7901 and AGS cell lines with stable knockdown of SNHG17 via shRNA targeting SNHG17 (Supplemental Figure 4D). Because SNHG17 was distributed in both the nuclear and cytosolic fractions of cells, SNHG17 distribution in the nuclear and cytosolic fractions of SNHG17-knockdown cells was also detected. As shown in Supplemental Figure 4E, SNHG17 levels in the nuclear and cytosolic fractions were both decreased. SNHG17 knockdown in gastric cancer cells resulted in a marked decrease in the formation of γ-H2AX foci upon H. pylori infection, as demonstrated by immunofluorescence and Western blotting (Figure 4 and Figure 5A). Moreover, SNHG17 knockdown also resulted in a marked decrease in ATM phosphorylation upon H. pylori infection, as demonstrated by immunofluorescence (Supplemental Figure 4F). In addition, knockdown of SNHG17 in SGC-7901 cells resulted in shorter comet tails than those in control cells in the neutral comet assay (Figure 5B).
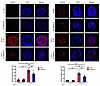 Figure 4
Figure 4SNHG17 knockdown decreased the accumulation of DSBs upon H. pylori infection. Representative photographs of γ-H2AX+ foci in the control and SNHG17-knockdown SGC-7901 (left) and AGS (right) cells infected with H. pylori. Original magnification, ×150 (left panels); ×180 (right panels). Data are represented as mean ± SEM. n = 5. **P < 0.01, ANOVA.
 Figure 5
Figure 5SNHG17 knockdown decreased the accumulation of DSBs upon H. pylori infection. (A) Western blot analysis of phosphorylation of γ-H2AX in SNHG17-knockdown SGC-7901 (top) and AGS (bottom) cells infected with H. pylori. Samples were assayed 3 times. (B) H. pylori–induced DNA damage in control and SNHG17-knockdown SGC-7901 cells, as measured by the comet assay. Scale bar: 10 μm. Data are represented as mean ± SEM. **P < 0.01, ANOVA. (C) SGC-7901–knockdown cells were infected with H. pylori for 6 hours, after which media were replaced with fresh media to allow the cells to recover. Western blotting analysis of phosphorylation of γ-H2AX was performed. Samples were assayed 3 times.
To analyze whether SNHG17 regulates DSB repair, we infected SGC-7901 with H. pylori for 6 hours and replaced the medium with fresh medium to allow the cells to recover. As shown in Figure 5C, the level of DNA damage gradually returned to the baseline level in the control cells after H. pylori infection, indicating efficient DSB repair; however, DNA repair was accelerated in SNHG17-knockdown cells.
Conversely, transiently overexpressing SNHG17 in normal gastric epithelial GES-1 cells (Supplemental Figure 5A) significantly enhanced the γ-H2AX levels (Figure 6, A and B). SNHG17 overexpression also resulted in a marked decrease in ATM phosphorylation upon H. pylori infection (Supplemental Figure 5B). Moreover, DNA repair was delayed in cells overexpressing SNHG17 (Figure 6C). Collectively, these data reveal the functional role of SNHG17 in DSB repair.
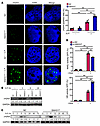 Figure 6
Figure 6SNHG17 expression increased the accumulation of DSBs upon H. pylori infection. (A) Representative photographs of γ-H2AX+ foci in control and SNHG17 overexpression GES-1 cells infected with H. pylori. Original magnification, ×150. Data are represented as mean ± SEM. n = 5. **P < 0.01, ANOVA. (B) Western blot analysis of phosphorylation of γ-H2AX in SNHG17-overexpressed GES-1 cells infected with H. pylori. Samples were assayed 3 times. (C) SNHG17-overexpressing GES-1 cells were infected with H. pylori for 6 hours, after which media were replaced with fresh media to allow the cells to recover. Western blot analysis of phosphorylation of γ-H2AX was performed. Samples were assayed 3 times. (D) GFP+ fraction of cells treated with SNHG17 siRNA indicated frequency of NHEJ repair or frequency of chromosomal DNA HR. I-SceI, an intron-encoded endonuclease. Data are represented as mean ± SEM. n = 3. **P < 0.01, ANOVA.
To further explore the mechanistic insights into SNHG17-mediated regulation of DSB repair during H. pylori infection, we evaluated the selection of the NHEJ or HR repair pathway by the transfection of SNHG17-knockdown cells with a system that enables flow cytometric analysis of the repair pathway selection at I-SceI–induced DNA breaks (30, 31). The results shown in Figure 6D revealed the SNHG17-dependent shift in the selected repair pathway; compared with control cells, SNHG17-knockdown cells showed decreased NHEJ repair and increased HR repair.
Nuclear SNHG17 directly interacted with NONO to regulate the NHEJ pathway of DSB repair. Because SNHG17 was located in the nucleus and cytoplasm simultaneously, we attempted to explore the possibility that SNHG17 functions by physically interacting with proteins or miRNAs. A biotinylated SNHG17 RNA pull-down assay and subsequent mass spectrometry (MS) analysis of the differentially displayed bands revealed that NONO was the main protein bound to SNHG17 (Supplemental Figure 6A and Supplemental Table 4). NONO is a characterized RNA- and DNA-binding nuclear factor. In addition, previous studies have shown that NONO can participate in the NHEJ pathway of DSB repair and stabilize broken DNA ends (32, 33).
The association between SNHG17 and NONO was further supported by immunoblotting the proteins captured in the SNHG17 RNA pull-down assay (Figure 7A). Ultraviolet light–crosslinking RNA immunoprecipitation (UV-RIP) assays with a NONO-specific antibody confirmed the direct interaction between SNHG17 and NONO (Figure 7B). Then, native RIP was performed to confirm binding between SHNG17 and NONO (Figure 7B); this binding was strengthened by H. pylori infection (Figure 7C). Moreover, the colocalization of SNHG17 and NONO was enhanced by H. pylori infection (Figure 7D).
 Figure 7
Figure 7Nuclear SNHG17 directly interacts with NONO. (A) RNA pull-down analysis of the binding of SNHG17 with NONO in total protein extracted from SGC-7901 cells infected with H. pylori. Samples were assayed 3 times. (B) NONO UV-RIP or native RIP followed by qPCR analysis of copurified RNA in SGC-7901 cells with H. pylori infection. Data are represented as mean ± SEM. n = 3. *P < 0.05, 2-tailed Student’s t test. (C) RNA pull-down analysis of the binding of SNHG17 with NONO in SGC-7901 cells infected with or without H. pylori. Samples were assayed 3 times. (D) Confocal microscopy images of SNHG17 stained with FISH probe (red) combined with immunofluorescence analysis of endogenous NONO (green) in SGC-7901 cells upon H. pylori infection. Nuclei were stained with DAPI (blue). Scale bar: 10 μm. Experiments were performed 3 times.
We then sought to determine the SNHG17-binding sites within the NONO protein. NONO consists of 2 tandem RRM domains (Supplemental Figure 6B), putatively responsible for protein-protein interactions with itself or for sequence-specific RNA binding. Glutathione-S-transferase (GST) pull-down assays using biotin-labeled SNHG17 were performed. Protein fragments containing either RRM1 or RRM2 or containing both RRM domains were used for an in vitro SNHG17-binding assay (Supplemental Figure 6C). We noted that both RRM1 and RRM2 played a role in the binding of NONO to SNHG17 (Figure 8A).
 Figure 8
Figure 8Nuclear SNHG17 directly interacts with NONO to regulate NHEJ pathway of DSB repair. (A) RNA pull-down analysis of the binding of the purified GST-NONO-RRM1, GST-NONO-RRM2, and GST–full-length NONO (NONO-FL) protein to in vitro–transcribed SNHG17. Samples were assayed more than 3 times. (B) A series of SNHG17 deletion mutants were transcribed in vitro and were used to perform RNA pull-down assays. Samples were assayed more than 3 times. (C) A NONO-binding site was predicated at location 716–720 nt of SNHG17. RNA pull-down analysis of the binding of NONO with SNHG17 probe or mutant SNHG17 probe. Samples were assayed more than 3 times. (D) CRISPR/Cas9 gene-editing–generated mutant SGC-7901 cells were validated by DNA sequencing. (E) NONO RIP followed by qPCR analysis of copurified RNA in CRISPR/Cas9 mutant SGC-7901 cells with or without H. pylori infection. (F) Distribution of primer pairs relative to the DSB created by I-SceI and quantification of NONO relative to the DSB by qRT-PCR. Data are represented as mean ± SEM. n = 3. *P < 0.05; **P < 0.01, 2-tailed Student’s t test (E); ANOVA (F).
To identify the binding domain on SNHG17 required for its interaction with NONO, we generated different truncation mutants of SNHG17. RNA pull-down assay suggested that the region of SNHG17 between nt 603 and 818 is required for its physical interaction with NONO (Figure 8B). We further used the RBPDB (database of RNA-binding protein specificities) database to predict and analyze the RNA-binding loci on SNHG17 and found a NONO-binding site (AGGGA) spanning nt 716–720 of SNHG17 (Figure 8C). After mutation of the predicted binding site, the binding capacity of SNHG17 to NONO was significantly weakened (Figure 8C), and SNHG17 was hypothesized to directly bind the NONO protein at the nt 716 to 720 site.
CRISPR/Cas9 gene-editing technology was used to target the NONO-binding site at nt 716 to 720 in order to generate mutant SGC-7901 cells and assess the function of the binding site. The homozygous SNHG17 mutant was validated by DNA sequencing (Figure 8D). Mutation of the nt 716 to 720 sequence did not alter the expression levels of either SNHG17 or the snoRNAs (Supplemental Figure 6D) but inhibited the binding of NONO to SNHG17 (Figure 8E).
NONO-bound SNHG17 might participate in regulating DSB repair and shift DSB repair toward NHEJ. Next, we investigated whether NONO binds to the ends of DSBs. ChIP qPCR was used to determine the position of NONO relative to DSB ends with a single I-SceI restriction site in SGC-7901 cells. Primers located at increasing distances from the DSB ends were used to evaluate the binding position of NONO (Figure 8F). NONO was detectable as close as 384 to 612 bp to the DSB, with higher enrichment than the IgG control. This localization resembled that of the NHEJ factor Ku80, as previously reported (34). However, when the expression of SNHG17 was inhibited, the enrichment of NONO at the 384 to 612 bp region was decreased (Figure 8F).
To further investigate the mechanism underlying the regulation of DSB by the binding of SNHG17 and NONO, we investigated whether SNHG17 and NONO regulate each other’s expression levels. As shown in Supplemental Figure 6, E and F, knockdown of neither SNHG17 nor NONO influenced the expression of the other molecule. The Ku70-Ku80 heterodimer binds to DSB ends as the initial step in DSB repair. NONO was reported to immunoprecipitate with Ku70 and Ku80, which participate in the NHEJ pathway of DSB repair (35). Thus, we examined whether SNHG17 interacts with Ku70 and Ku80 and showed that SNHG17 did not immunoprecipitate with Ku70 or Ku80 (data not shown). This observation motivated us to propose that the physical association of SNHG17 and NONO might affect the interaction between NONO and Ku70/Ku80 upon H. pylori infection.
Cytoplasmic SNHG17 interacted with miR-3909 to regulate the HR pathway of DSB repair. Our evidence revealed that NHEJ activity was decreased and HR activity was increased in SNHG17-knockdown cells and that nuclear SNHG17 directly interacted with NONO to regulate the NHEJ pathway of DSB repair. We then investigated the mechanism by which SNHG17 regulates the HR pathway. As previously reported (6), expression of the central HR protein Rad51 was decreased upon H. pylori infection. We then investigated Rad51 expression in SNHG17-knockdown cells and showed that H. pylori infection decreased Rad51 expression and SNHG17 knockdown enhanced Rad51 expression, as shown in Figure 9A. The mechanism by which the Rad51 increase was induced by SNHG17 knockdown was further investigated. The Rad51 mRNA levels remained unaltered in SNHG17-knockdown cells (Supplemental Figure 7A), and SNHG17 knockdown did not influence the degradation of Rad51 mRNA in the presence of actinomycin D (Act D) (Supplemental Figure 7B). Western blot analysis after the addition of cycloheximide (CHX) to block translation indicated that SNHG17 knockdown did not affect the regulation of Rad51 protein translation (Supplemental Figure 7C). Taken together, these results indicate that the altered Rad51 level might be due to the regulation of Rad51 protein stability.
 Figure 9
Figure 9SNHG17 interacts with miR-3909. (A) Western blot analysis of Rad51 expression upon H. pylori with SNHG17 knockdown. Samples were assayed more than 3 times. (B) Western blot analysis of Rad51 in cells expressing the indicated shRNA in the presence or absence of MG-132 treatment. Samples were assayed more than 3 times. (C) Bioinformatics predicted miR-3909 and RING1-binding sites in SNHG17 sequence. Partial sequences of SNHG17 and RING1 3′ UTR are shown. (D) qRT-PCR analysis of the expression levels of pri-miR-3909, pre-miR-3909, and mature miR-3909 in SGC-7901 cells overexpressing SNHG17. (E) qRT-PCR analysis for SNHG17 expression in SGC-7901 cells transfected for 24 hours with miR-NC, miR-3909 mimics, miR-NC inhibitor, and miR-3909 inhibitor. Data are presented as mean ± SEM. n = 3. *P < 0.05, 2-tailed Student’s t test.
RING1 was reported to induce Rad51 degradation through the ubiquitin-proteasome pathway (UPP) (36). We found that MG-132 treatment increased the Rad51 level (Figure 9B), indicating that Rad51 is a target for degradation by the UPP. Recent accumulating evidence has shown that lncRNAs are regulated by miRNAs through mutual combination as competing endogenous RNAs (ceRNAs). The starBase, version 2.0 (http://starbase.sysu.edu.cn/index.php), bioinformatics website predicted that SNHG17 could bind to miR-3909 (Figure 9C), an miRNA-targeting RING1. Based on this prediction, we hypothesized that SNHG17 might act as a sponge upon H. pylori infection. Correspondingly, miR-3909 expression was demonstrated to decrease upon H. pylori infection (Supplemental Figure 7D).
miR-3909 expression was significantly decreased in cells overexpressing SNHG17 compared with that in control cells (Figure 9D). However, SNHG17 affected the expression of neither pri-miR-3909 nor pre-miR-3909, indicating that SNHG17-mediated regulation of miR-3909 expression likely occurs through a posttranscriptional mechanism. The effect of miR-3909 on SNHG17 expression was further analyzed by qRT-PCR. As shown in Figure 9E, the miR-3909 mimic significantly reduced SNHG17 expression; conversely, the miR-3909 inhibitor markedly increased SNHG17 expression, demonstrating reciprocal repression between SNHG17 and miR-3909.
To determine whether miR-3909 recognized the predicted target site within SNHG17, we constructed luciferase vectors expressing WT and mutant SNHG17 (in which the miR-3909–binding motif was mutated). Using dual luciferase assays, we found that transfection of the WT SNHG17 vector, but not the mutant SNHG17 vector, with the miR-3909 mimic significantly decreased luciferase activity (Figure 10A), indicating that miR-3909 can bind to SNHG17 through an miRNA recognition site.
 Figure 10
Figure 10SNHG17 interacts with miR-3909 to regulate HR pathway of DSB repair. (A) Relative firefly luciferase activity derived from psiCHECK2-SNHG17-WT or psiCHECK2-SNHG17-mut plasmid cotransfected with miR-3909 mimic into HEK293 cells for 48 hours. (B) Relative firefly luciferase activity derived from psiCHECK2-RING1-WT or psiCHECK2-RING1-mut plasmid cotransfected with miR-3909 mimic into HEK293 cells for 48 hours. (C) RIP with an anti-Ago2 antibody was used to assess endogenous Ago2 binding to RNA; IgG was used as the control. Levels of SNHG17 and miR-3909 were determined by qRT–PCR. (D) Western blot analysis was performed to test Rad51 protein expression after SGC-7901 cells were transfected with miR-3909 mimic or inhibitor for 48 hours. (E) Western blot analysis was performed to test RING 1 and Rad51 protein abundance after SGC-7901 cells were transfected with miR-3909 mimics and/or the SNHG17 overexpression vector. Samples were assayed more than 3 times. Data are represented as mean ± SEM. n = 3. *P < 0.05; **P < 0.01, 2-tailed Student’s test.
Next, we performed reporter assays with a luciferase plasmid harboring the 3′ UTR sequence of RING1, which contains the predicted miR-3909–binding site. Furthermore, we constructed mutant reporter vectors containing a mutation in the miR-3909–binding sites (RING1-3′ UTR–Mut). These plasmids were cotransfected into HEK293T cells with the miR-3909 mimic. Reporter assays in 293T cells revealed miR-3909–dependent repression of the RING1 3′ UTR. The mutation abolished repression by miR-3909, indicating that miR-3909 specifically targeted the binding sites in the RING1 3′ UTR (Figure 10B).
To determine whether SNHG17 was included in miRNA-containing RNA-induced silencing complexes (RISCs), RIP experiments were performed in SGC-7901 cell extracts using an anti-Ago2 antibody. SNHG17 and miR-3909 were enriched in miRNA-containing RISCs that included Ago2 relative to their levels in control IgG immunoprecipitates, suggesting that the Ago2 protein bound directly to both SNHG17 and miR-3909 in SGC-7901 cells (Figure 10C). Consistent with the results of the above assay, Rad51 protein expression in SGC-7901 cells was decreased in the presence of the miR-3909 inhibitor (Figure 10D). Conversely, Rad51 expression levels increased after treatment with the miR-3909 mimics (Figure 10D).
To verify that SNHG17 interacted with miR-3909 as a ceRNA to regulate the activity of the RING1 3′ UTR, we examined the RING1 protein levels by Western blot analysis. Upregulation of SNHG17 promoted RING1 protein expression and thus decreased Rad51 protein expression, which was rescued by overexpression of miR-3909 (Figure 10E). These results strongly indicate that SNHG17 regulates RING1/Rad51 expression in a miR-3909–dependent manner.
SNHG17 knockdown regulated genome rearrangements in gastric cancer cells. Our results demonstrated that the nt 702 to 708 sequence of SNHG17 was complementary to the seed sequence of miR-3909 and that the nt 716 to 720 region was the NONO-binding site. The binding sites for miR-3909 and NONO were very close to each other, indicating that miR-3909 and NONO cannot bind to SNHG17 simultaneously. To further clarify the mechanism by which SNHG17 shifts DSB repair from the HR to the NHEJ pathway, Rad51 expression was investigated in SGC-7901 cells expressing SNHG17 with a mutated nt 716 to 720 sequence. Rad51 inhibition induced by H. pylori infection was not altered by the SNHG17 716 to 720 nt mutation (Figure 11A). In addition, the formation of γ-H2AX foci was increased in SNHG17 mutant–expressing SGC-7901 cells, as demonstrated by the immunofluorescence results (Figure 11B). Further, the expression of p-ATM is also increased in SGC-7901 cells with SNHG17 mutation (Supplemental Figure 8A). Moreover, the results of the comet assay showed that SNHG17 mutant–expressing SGC-7901 cells exhibited longer comet tails than control cells infected with H. pylori (Supplemental Figure 8B). The inhibition of DSB repair might be due to a deficiency in NONO binding. These results provide convincing evidence that the distinct mechanisms by which SNHG17 regulates the DSB pathway are spatially independent.
 Figure 11
Figure 11SNHG17 NONO–binding site mutation inhibited DSB repair. (A) Western blot analysis of Rad51 expression in SNHG17 mutant SGC-7901 cells infected with H. pylori. Samples were assayed more than 3 times. (B) Representative photographs of γ-H2AX–positive foci in SNHG17 mutant SGC-7901 cells infected with H. pylori. Original magnification, ×40. Experiment was conducted 3 times. Data are represented as mean ± SEM. n = 5. **P < 0.01, 2-tailed Student’s t test.
Excessive NHEJ activity leads to frequent chromosome aberrations and genome rearrangements that might contribute to tumor progression (37). To determine whether the SNHG17-driven upregulation of NHEJ activity renders the gastric cancer cell line more prone to genomic instability, a chronic H. pylori infection model was constructed; chronic H. pylori infection is a strong risk factor for stomach cancer. To gain an overview of the landscape of genetic alterations that accumulated in the gastric cancer cell line SGC-7901 with chronic H. pylori infection, we performed whole-genome sequencing of the constructed chronic infection cell model. Short insertions and deletions (indels), chromosomal structural variations (SVs), fusion genes, and copy number variants (CNVs) were identified. We found that the genomes of SGC-7901 cells infected with H. pylori harbored significantly lower numbers of CNVs and higher numbers of indels, SVs, and fusion genes than did the genomes of SNHG17-knockdown SGC-7901 cells infected with H. pylori (Figure 12A).
 Figure 12
Figure 12SNHG17 knockdown regulated genome rearrangements in gastric cancer cells. (A) Short indels, chromosomal SVs, CNVs, and fusion genes detected by whole genome sequencing of the constructed chronic H. pylori infection SGC-7901 cell model. (B) Growth of SNHG17-knockdown SGC-7901 cells injected subcutaneously into nude mice (subcutaneous xenografts). n = 7. (C) Growth of chronic infection SGC-7901 cells injected subcutaneously into nude mice (subcutaneous xenografts). n = 10. Data are represented as mean ± SEM. **P < 0.01, 2-tailed Student’s t test.
The sequencing data showed that SNHG17 knockdown regulated chromosomal aberrations and genome rearrangements, a hallmark of cancer cells (38, 39). Thus, the growth of H. pylori–infected SGC-7901 cells was evaluated in vivo to assess whether SNHG17 could contribute to tumor progression. The growth of SNHG17-knockdown SGC-7901 cells in subcutaneous xenografts was not altered compared with that of control cells (Figure 12B). However, the growth of SNHG17-knockdown SGC-7901 cells with chronic H. pylori infection in subcutaneous xenografts was significantly inhibited compared with that of control cells (Figure 12C).
In summary, our study revealed an lncRNA, SNHG17, associated with H. pylori–infected gastric cancer that functions as a regulatory lncRNA to control the selection of the DSB repair pathway by facilitating the formation of the NHEJ DSB repair complex through interacting with NONO and by inhibiting HR activity through interacting with miR-3909 (Figure 13). Therefore, the results of this study show that SNHG17 is a lncRNA that affects gastric cancer cell genomic stability and might also be a biomarker for the early diagnosis and treatment of gastric cancer.
 Figure 13
Figure 13Proposed working model for SNHG17 regulation of genomic instability upon H. pylori infection. In brief, dysregulation of SNHG17/NONO and SNHG17/miR-393909/RING1/Rad51 pathways upon H. pylori infection drives the cells more prone to genome rearrangements, thus promoting tumorigenesis in gastric cancer.
-
Discussion
The results of this study revealed that SNHG17, identified by a coexpression network based on microarray screening, is associated with H. pylori–induced DSB repair and thus might promote gastric cancer progression. According to previous studies, core gene networks and hub genes associated with progression of disease were picked out by coexpression modules (40–42). In our study, differentially expressed lncRNA and mRNA in H. pylori–infected GES-1 cells at different time points were chosen for calculating pairwise Pearson’s correlation coefficient. In fact, we also reported SNHG17 upregulation in gastric cancer in a previous publication–based panel of screened snoRNA host genes (SNHGs)(43). These results suggest a critical role for snoRNAs or their host genes in cancer progression. In addition to our study on SNHG5 and SNHG17 in gastric cancer, other studies on the functions of SNHGs in cancer progression have also been published (44). We found that the basal expression levels of the majority of SNHGs were high. An absolute copy number analysis revealed that SNHG17 was present at a high level of 180 molecules per uninfected GES-1 cell and increased to approximately 1450 copies per cell upon H. pylori infection for 24 hours (data not shown), a higher abundance than that of other lncRNAs (45), suggesting that alterations in SNHGs might play important roles in cell behavior. Moreover, SNHG17 intronic snoRNAs were upregulated upon H. pylori infection and in gastric cancer tissues, indicating that these molecules might also be regulators in gastric cancer. However, the function of these snoRNAs needs further investigation in the future.
Upon bacterial infection, a major challenge for host cells is the maintenance of genomic integrity. H. pylori has been found to cause several types of DNA damage, including single-strand breaks and DSBs (6, 15, 17). In addition, H. pylori increases the mutation rate and chromosomal instability of host cells (46, 47). The results of high-throughput genomic analyses have shown that H. pylori causes a specific pattern of DNA damage, targeting transcribed and telomere-proximal regions correlating with the previously described genomic alterations typical of gastric cancer (17). These findings strongly support and emphasize the role of H. pylori–induced DNA damage in gastric carcinogenesis. However, the mechanisms of H. pylori–induced host genomic instabilities remain poorly understood. As a result of DNA damage, H. pylori infection leads to the activation of DNA damage repair. Under conditions of chronic infection, DNA damage repair appears to be insufficient for complete repair of DNA damage, thus resulting in mutations and chromosomal instability. H. pylori has been demonstrated to systematically reduce or modulate the activity of multiple cellular pathways involved in DNA damage repair, such as base excision repair (BER), mismatch repair (MMR), NHEJ, and HR (48). The latter 2 repair pathways (NHEJ and HR) are responsible for repairing DSBs, which are the most severe type of DNA damage. A shift in the repair of DNA DSBs from HR to NHEJ during H. pylori infection, possibly leading to a more error-prone repair process and thereby contributing to genetic instability in infected cells, was hypothesized (17). Indeed, our results showed that the overexpression of SNHG17 and the recruitment of NONO along with the role of cytoplasmic SNHG17 as a decoy for miR-3909, which regulates Rad51 expression, shift the DSB repair balance from HR toward NHEJ, supporting the hypothesis that H. pylori–induced DSBs are preferentially repaired by the NHEJ pathway (49). In our study, the shift in DNA repair from the HR toward the NHEJ pathway led to enhanced γ-H2AX foci formation, indicating a delay in DSB repair. This delay might be explained by the observation that the decrease in HR pathway activity seemed to be greater than the increase in NHEJ pathway activity or possibly by the increased abundance of SNHG17 in the cytoplasm relative to that in the nucleus upon H. pylori infection.
Our results demonstrated that the nt 702 to 708 sequence of SNHG17 was complementary to the seed sequence of miR-3909 and that the nt 716 to 720 region was the NONO-binding site. These 2 binding sites were very close to each other, indicating that miR-3909 and NONO cannot bind to SNHG17 simultaneously. In this study, we applied CRISPR/Cas9 gene editing technology to mutate the NONO-binding site in SNHG17 in SGC-7901 cells to clarify the function and mechanism of SNHG17. In the mutant cells, only the NONO-binding site was edited by CRISPR/Cas9; the miR-3909–binding site was not affected. The inhibition of DSB repair in the mutant cells should thus be due to the deficiency in NONO binding and unaltered Rad51 levels. The results provided convincing evidence that the distinct mechanisms by which SNHG17 regulates the HR and NHEJ pathways simultaneously are spatially independent.
The SNHG17 target mutation constructed in this study is site-directed mutation. Single-stranded oligonucleotides (ssODNs) should be used as HR template, and CRISPR/Cas9 be used to construct monoclonal cell lines carrying mutation, but the editing efficiency is limited. Considering that only the 5-nt sequence needs to be deleted, random deletion of DNA double strand by carrying Cas9 to the target site by gRNA was produced. A large number of clones were screened to obtain biallelic deletion clones for the target site, and we have also obtained some clones with partial deletion of the target site in this process. These clones could also be used as research tools for further studying. At the same time, we also used the HR method to screen the clones with target site deletion and obtained the target clones with this method as well.
DSBs originate when both DNA strands are broken at the same position or in sufficiently close proximity to allow the physical dissociation of the double helix. In addition to loss of genetic information, DSBs can lead to fragmentation, loss, or rearrangement of chromosomes. Deregulated DNA repair can result in chromosomal translocation, genomic rearrangement, and an increased mutation rate, which might provide survival advantages to cancer cells (50–52). Clearly, maintaining the proper balance of DSB repair is critical for preventing pathologic chromosomal rearrangements and subsequent tumor development. Moreover, knowledge of defects in DNA repair pathways provides a valuable opportunity for exploring the potential of these pathways as therapeutic targets in cancer. Some inhibitors designed as cancer therapeutics specifically targeting DSB repair proteins have been reported (52, 53). However, DSB repair pathway selection and the associated factors remain poorly defined, especially regarding the mechanism by which H. pylori–induced DSB repair accounts for gastric cancer development. To elucidate the effect of abnormal DSB repair on tumor growth, we established a modified chronic infection model based on a previous report (46) to imitate in vivo infection conditions. Furthermore, a subcutaneous transplanted tumor model in nude mice was used to eliminate other possible influencing factors induced by H. pylori infection, such as cytokine variations, immune cells, and epithelial cell interaction. We showed that the growth of SNHG17-knockdown SGC-7901 cells with chronic H. pylori infection in subcutaneous xenografts was inhibited compared with that of control cells. Notably, sequencing results have shown that abnormal expression of SNHG17 has a marked effect on chromosome stability. These results provide clear evidence and mechanistic insight into the means by which SNHG17 regulates DSB repair to promote gastric cancer development.
When homology is not used to ensure that molecules are rejoined in the correct positions, there is some probability that genetic change will result (54). Although HR provides vital repair mechanisms, it also can cause chromosomal structural change, including CNV (55). Segmental duplication–related nonallelic HR is an important mechanistic driver of SV hotspot formation (56). Nonallelic HR is an important evolutionary mechanism for the creation of pseudogenes and the creation of novel genes via fusion (57). NHEJ has been shown to underlie the rearrangements causing genomic disorders (58). NHEJ is capable of generating deletions with microhomology at the break points. In the study of Kidd et al. (59), an inferred 20.6% of the SVs were due to NHEJ, 36% due to NHEJ plus microhomology-mediated end joining (MMEJ), and 10.0% due to NHEJ plus fork stalling and template switching (FoSTeS). NHEJ is also important for chromosome translocations, which create neomorphic fusion genes occurring in both lymphoid malignancies and solid tumors. Many translocations arise as a consequence of “classical” or “alternative” pathways of NHEJ (60). The work of Shaw et al. has shown that androgen stimulation increases the recruitment of several proteins involved in the error-prone NHEJ pathway to the TMPRSS2-ERG fusion break points and that inhibiting various components of the NHEJ pathway decreased the generation of the TMPRSS2-ERG gene fusions (58). The error-prone NHEJ pathway often results in small indels, while the HR pathway results in precise repair with a homologous chromosome or an exogenous donor template (61).
H. pylori infection will induce DNA damage and regulate DNA repair. Several studies have shown that H. pylori infection induces genetic instability of both the nuclear and the mitochondrial DNA (46). In the work of Bibi et al., copy number gains were more frequent than losses throughout all GC samples compared with normal tissue samples. DNA copy number gains at 1p36.32, 2p11.1, 4q23–q25, 5p12-p11, 6p21.33, 9q12-q21.11, 12q11–q12, 14q32.33, 16p13.3, 17p13.1, 17q25.3, and 19q13.32 and losses at 1p36.23, 1p36.32, 1p32.1, 3q25.2, 6p21.33, 8p11.22, and 16q24.2 may be common in GC. Of these, 60% of the samples were positive for H. pylori infection (62). The API2-MALT1 fusion gene was originally identified from a t(11;18)(q21;q21) translocation, a specific chromosomal abnormality that is found in mucosa-associated lymphoid tissue (MALT) lymphoma. Gastric MALT lymphomas positive for the API2-MALT1 fusion gene do not respond to H. pylori eradication therapy (63). Studies by Machado et al. show that the majority of mutational events in H. pylori–infected cells were transitions (76.9%). Transversions were 7.7%, insertions were 11.5%, and deletions were 3.8% (64). In our study, because the chronic infection model was based on SGC-7901 cells, the sequencing data were compared with a reference genome for data analysis. Then, SGC-7901 cells were used as WT, and the sequencing data of other groups were compared with SGC-7901 to quantify the differences of variation. The results indicated that the genomes of SGC-7901 cells infected with H. pylori harbored substantially lower numbers of CNVs and higher numbers of indels, SVs, and fusion genes than did the genomes of SNHG17-knockdown SGC-7901 cells infected with H. pylori. Indeed, these results supported that SNHG17 shifted the H. pylori–induced DSB repair balance from HR to NHEJ.
In summary, SNHG17 was markedly upregulated in H. pylori–positive atrophic gastritis and gastric cancer. Moreover, SNHG17 is an lncRNA that affects the genomic stability of gastric cancer cells, thus contributing to the progression of H. pylori–induced gastric cancer. This assertion is supported by data implicating aberrant SNHG17 expression in gastric cancer and indicating that SNHG17 is a potential biomarker for the early diagnosis and treatment of gastric cancer. Moreover, the results of this study provide a basis for the future understanding of the response of gastric epithelial cells to H. pylori and the contributing effect of this response to gastric cancer development. Upregulation of SNHG17 by H. pylori infection appears to explain at least one of the missing links between cancer and inflammation.
-
Methods
Cell lines and siRNAs. The sources of the gastric cancer cell lines and the normal gastric immortalized epithelial cell line GES-1 have been described previously (39). HEK293T (ATCC) and MGC-803 cells were cultured in DMEM (Gibco, Thermo Fisher Scientific), and all other cell lines were cultured in RPMI 1640 medium (Gibco, Thermo Fisher Scientific) supplemented with 10% FBS (Gibco, Thermo Fisher Scientific). All cultures were maintained at 37°C and 5% CO2. Cells were transiently transfected using Lipofectamine 2000 (Thermo Scientific). SNHG17 siRNAs, NONO siRNAs, and control siRNAs were obtained from Invitrogen, Thermo Fisher Scientific.
Bacterial strains and infection. H. pylori strain NCTC11637 was purchased from ATCC. H. pylori was inoculated onto Columbia blood agar base medium containing 10% defibrinated sheep blood and incubated at 37°C in a microaerobic environment (5% O2, 10% CO2, 85% N2). After culture for 3 days, H. pylori was harvested.
Clinically isolated strains were provided by the Fourth Affiliated Hospital of Hebei Medical University. Antral gastric biopsy samples obtained from microscopic examination were inoculated on Colombia blood agar base medium containing 5% defibrinated sheep blood and incubated at 37°C in a microaerobic environment (5% O2, 10% CO2, 85% N2). After culture for 3 to 7 days, single colonies were picked and isolated. After staining and biochemical analysis, all strains that were gram negative and urease, catalase, and oxidase positive were identified as H. pylori. Then the H. pylori strains were inoculated into brain heart infusion (BHI) broth containing 30% glycerin and cryopreserved at –70°C. The genotypes of clinical strain 1 and clinical strain 2 were cagA+, cagE+, and vacA s1m1; and cagA+, cagE+, and vacA s2m2, respectively.
For the acute infection model, cells grown to 80% confluence were washed with PBS and incubated in antibiotic-free medium. Mature colonies (72 hours) of H. pylori were collected, and the bacterial cells were added to the human cells at the indicated MOI. For the chronic infection model, SGC-7901 cells grown to 30% confluence were treated similarly to the cells in the acute infection model. Every 72 hours, the cells were washed with PBS, and fresh antibiotic-free medium and bacterial cells were added at an MOI of 10. This process was repeated 10 times to obtain chronically infected gastric cancer cells.
“Dead” H. pylori were obtained by incubation in 100% ethanol. Bacteria cocultured with host cells were killed with penicillin/streptomycin combined with 15 μg/mL metronidazole, 30 μg/mL tetracycline hydrochloride (both from MilliporeSigma), and 4 μg/mL bismuth subcitrate (Selleck).
Patients and specimens. All clinical tissue samples from patients were obtained from the Fourth Affiliated Hospital of Hebei Medical University and the First Affiliated Hospital of Harbin Medical University between 2013 and 2018. H. pylori–infected gastric epithelial tissues and premalignant tissues with intestinal metaplasia or dysplasia were obtained endoscopically. Histological specimens were diagnosed by 2 trained pathologists in a blinded manner. Gastric cancer tissues originating from patients with primary gastric cancer with tumor stages I to IV for whom surgery was the initial treatment approach were included. The pathologic stage was assessed according to the revised international system (Gastric Cancer, Version 3.2016, NCCN Clinical Practice Guidelines in Oncology) (65). H. pylori infection status was confirmed by a C-14 urea breath test and/or histological testing. Patients were considered positive for H. pylori infection when at least one of these examinations yielded positive results and were considered H. pylori negative if all test results were negative.
lncRNA expression microarray analysis. The lncRNA and mRNA expression microarray was performed by KangCheng Biotechnology Co. Total RNA was extracted from cells and biopsy tissues using TRIzol reagent (Invitrogen, Thermo Fisher Scientific). The quantity of extracted RNA was determined using a NanoDrop spectrophotometer (Thermo Fisher Scientific). The quality of the total RNA was further analyzed by electrophoresis to assess integrity or contamination. Purified RNA was used to synthesize double-stranded cDNA, which was labeled (Agilent Technologies, 5190-0442) and hybridized to the human lncRNA/mRNA Array, version 2.0 (8 × 60 K; Arraystar). The microarray was then scanned using an Agilent G2505B Microarray Scanner (Agilent Technologies). Raw data were analyzed with Feature Extraction software (version 11.0.1.1, Agilent Technologies). Quartile normalization and data analysis were conducted using the GeneSpring GX, version 11.5.1, software package (Agilent Technologies). Samples were grouped and then compared. Differentially expressed lncRNAs were subjected to hierarchical clustering based on Cluster/TreeView software from Stanford University. Microarray data were deposited in the NCBI’s GEO database (GSE111762, GSE111763). Moreover, whole genome sequencing of SGC-7901 cells chronically infected with H. pylori and subsequent bioinformatic analysis were performed by Annoroad Genome Co. The sequencing data were deposited in the NCBI’s Sequence Read Archive (SRA BioProject SRR10448203, SRR7600350, SRR10447757, and SRR10478832). The sequencing data were compared with a reference genome for data analysis. Then SGC-7901 cells were used as WT, and the sequencing data of other groups were compared with SGC-7901 to quantify the differences of variation.
Coexpression network. Gene coexpression networks were constructed according to the normalized signal intensity of specifically expressed genes, performed for us by CNkingbio Co. For each pair of genes, Pearson’s correlation coefficient was calculated by the R function cor.test, and the significantly correlated pairs, with cutoff defined as 0.9999, were chosen to construct the network. Degree centrality was defined as the number of links between 2 nodes. Visualization of the network was built by the software Cytoscape, version 3.6.0.
Biotin-RNA pull-down assay and deletion mapping. pGEM-3zf(+)-SNHG17 and pGEM-3zf(+)-SNHG17-antisense plasmids were linearized by restriction enzyme digestion for use as templates for the transcription of SNHG17 and SNHG17 antisense sequences and as template DNAs for various fragments of SNHG17. Sequences were amplified from GES-1 RNA by RT-PCR using primers listed in the Supplemental Methods. Biotin-labeled RNAs were transcribed in vitro using Biotin RNA Labeling Mix (Roche), treated with RNAse-free DNase I (TaKaRa), and purified with an RNeasy Mini Kit (Roche). Biotinylated RNA in RNA structure buffer (10 mM Tris [pH 7], 0.1 M KCl, and 10 mM MgCl2) was heated to 98°C for 2 minutes, placed on ice for 5 minutes, and then left at room temperature for 25 minutes to allow proper secondary structure formation. Total protein lysates of SGC-7901 cells infected with H. pylori (MOI, 200:1) were mixed with the biotinylated RNA and incubated at 30°C. Streptavidin agarose beads (GE Healthcare) were added to each binding reaction and further incubated at room temperature with rotation. RNA–protein bead complexes were washed with binding buffer, dissolved in nuclease-free water and PBS, and boiled in SDS-PAGE loading buffer for 10 minutes. RNA affinity capture samples were subjected to 12% SDS-PAGE and visualized by silver staining or immunoblotting. Protein bands were excised and identified by in-gel trypsin digestion followed by liquid chromatography–tandem MS (LC-MS/MS) (nanoLC-LTQ-Orbitrap XL, Thermo Fisher Scientific) or hybridization with anti-NONO antibody (Abcam, catalog ab70335).
RIP and UV-RIP. RIP was performed using a Magna RIP RNA-Binding Protein Immunoprecipitation Kit (MilliporeSigma) according to the manufacturer’s instructions, with slight modifications. Briefly, antibodies directed against NONO (Abcam, catalog ab70335) or Ago2 (Abcam, catalog ab32381) were used. Coprecipitated RNAs were adsorbed to magnetic beads and detected by reverse-transcription PCR. Total RNA (input controls) and isotype control rabbit IgG antibody were used simultaneously to demonstrate that the detected signals arose from RNAs specifically binding to NONO or other proteins. UV-RIP was performed according to the RIP protocol with the following modifications: cells were incubated for 12 hours with 100 mM 4-thiouridine (4-SU) and were then cross-linked using 365 nm UV light at a dose of 400 mJ/cm2. The primers used for RIP-qRT-PCR analysis are listed in Supplemental Table 5.
Comet assay. Cells were mixed with 0.5% low–melting point agarose. The mixture was spread on slides precoated with normal agarose (1% in PBS) at 37°C and cooled to solidify using ice packs for 5 minutes. After the agarose was solidified, cells were immersed in lysis solution (2.5 M NaCl, 100 mM EDTA, 10 mM Tris, 1% Triton X-100, and 10% DMSO [pH 10]) for 1 hour at 4°C. Slides were placed in a gel electrophoresis apparatus containing 300 mM NaOH and 10 mM Na-EDTA (pH 13) for 30 minutes. After electrophoresis, slides were rinsed 3 times with neutralizing buffer (0.4 M Tris, pH 7.5) for at least 5 minutes each, dehydrated in absolute ethanol at 4°C, and allowed to dry. Cells were stained with 20 μg/mL propidium iodide (PI) (MilliporeSigma). Images were then captured using a fluorescence microscope. Quantification of tail DNA was performed with CASP software.
GST pull-down. cDNA of NONO or its domains was cloned into the pGEX-6P-1 vector (GE Healthcare) and expressed as a GST fusion protein in Escherichia coli. The expressed proteins were purified by Glutathione Sepharose 4B. Probes for the full-length SNHG17 sequence were transcribed in vitro from a plasmid containing the T7 promoter by T7 polymerase using Biotin RNA Labeling Mix (Roche), treated with RNAse-free DNase I (TaKaRa), and purified with the RNeasy Mini Kit (Roche). Biotinylated RNA in RNA structure buffer (10 mM Tris [pH 7], 0.1 M KCl, and 10 mM MgCl2) was heated to 98°C for 2 minutes, placed on ice for 5 minutes, and then left at room temperature for 25 minutes to allow proper secondary structure formation. Purified GST fusion proteins were mixed with biotinylated RNA and incubated at 30°C. Streptavidin agarose beads (GE Healthcare) were added to each reaction and further incubated at room temperature with rotation. RNA–protein bead complexes were washed with binding buffer, dissolved in nuclease-free water and PBS, and boiled in SDS-PAGE loading buffer for 10 minutes. RNA affinity capture samples were subjected to 12% SDS-PAGE and transferred to PVDF membranes. Membranes were probed with an anti-GST antibody (Cell Signaling Technology, catalog 2624) and secondary antibodies, followed by visualization with ECL (Amersham Biosciences).
CRISPR/Cas9 gene targeting, selection, and removal of selection markers. To create cell lines expressing SNHG17 with deletion of the NONO-binding site, HR with donor DNA based on CRISPR/Cas9 editing was not used in this study because of the low efficiency of this approach. Two guide RNAs were cloned into the PX458 vector and individually transfected into SGC-7901 cells. GFP-expressing colonies were then sorted by flow cytometry (BD FACSAria III Cell Sorter, Beckman Coulter) to enrich for targeted cells. Clones were plated in 96-well plates via a limited dilution method at a density of 0.8 cells per well. Deletion mutants were identified by sequencing.
Animal experiments. Six- to eight-week-old male BALB/c nude (Weitong Lihua) mice were used to determine SNHG17 effects in vivo. Nude mice were injected subcutaneously in the right armpit with 1 × 106 SGC-7901 cells. Tumor size was measured every 3 or 5 days until the tumors attained a volume of approximately 1500 mm3.
TCGA data download and analysis. RNA-Seq data of gastric cancer tissues and paracancerous tissues were searched in the TCGA cancer database, and the RNA-Seq data were downloaded using the gdc-client.exe tool and the manifest file provided in the TCGA database. The download command is gdc-client.exe download –m manifest filename. Then we used Perl language to write scripts for the RNA-Seq data format and ran the Perl script on a Linux server to extract and integrate RNA-Seq data in order to obtain the RNA-Seq matrix of gastric cancer and paracancerous tissues. At the same time, we downloaded the Homo_sapiens.GRCh38.88.chr.gtf human genome annotation file in the ENSEMBLE database and used the human genome annotation file to convert the ensemble id in the matrix into the gene ID and then obtained the RNA-Seq matrix containing the gene ID. The R language script was used to analyze the differences in gene expression of RNA-Seq data, and the logFC, logCPM, P values, and FDR values of all genes were obtained. Finally all the genes with differential expression in gastric cancer were obtained.
Statistics. Statistical analyses were performed using SPSS, version 13.0, software. Results are expressed as mean ± SEM and were analyzed by 2-tailed Student’s t test (unpaired) or 1-way ANOVA followed by Bonferroni’s test. The χ2 test was used to analyze categorical variables. P values of less than 0.05 were considered statistically significant.
Study approval. Informed consent for the use of the samples was obtained from all patients, and the study was approved by the ethics committee of the Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences, and the ethical committees of the Fourth Affiliated Hospital of Hebei Medical University and the First Affiliated Hospital of Harbin Medical University. All mouse experiments were carried out in accordance with institutional animal guidelines and were approved by the IACUC of the Center for Experimental Animal Research (Beijing, China).
-
Author contributions
TH and J Shi were involved in the overall design of experiments and interpretation of results. J Sun was involved in interpretation of some results. TH designed and prepared constructs. AZ and SZ performed the bioinformatics analysis. TH performed ISH, immunofluorescence staining, and experiments for RNA-pull-down and RIP assays. XJ performed CRISPR/Cas9 studies. JB and HG performed experiments involving human specimens. RM performed some WB experiments. BZ and LZ supervised the experiments involving human materials.
-
Acknowledgments
We thank Yabin Guo (Medical Research Center, Sun Yat-Sen Memorial Hospital, Sun Yat-Sen University, Guangzhou, China) for the suggestion on bioinformatic analysis. This work was partially supported by the Natural Science Foundation of China (81572755, 31771535 to J Shi and the CAMS Initiative for Innovative Medicine (CAMS-I2M 2016-I2M-1-001 to J Shi).
Address correspondence to: Juan Shi, Institute of Basic Medical Sciences, 5 Dong Dan San Tiao, Beijing 100005, China. Phone: 86.10.69156430; Email: [email protected], [email protected].
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Copyright: © 2020, American Society for Clinical Investigation.
Reference information: J Clin Invest. 2020;130(7):3901–3918.https://doi.org/10.1172/JCI125581.
-
References
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
- International Agency for Research on Cancer (IARC) Working Group on the Evaluation of Carcinogenic Risk to Humans. Infection with Helicobacter pylori. In: IARC Working Group on the Evaluation of Carcinogenic Risk to Humans, eds. Schistosomes, Liver Flukes and Helicobacter pylori: Views and Expert Opinions of an IARC Working Group on the Evaluation of Carcinogenic Risks to Humans, which Met in Lyon, 7-14 June 1994. Geneva, Switzerland: WHO; 1994:177–240.
- Wroblewski LE, Peek RM, Wilson KT. Helicobacter pylori and gastric cancer: factors that modulate disease risk. Clin Microbiol Rev. 2010;23(4):713–739.
- Hanada K, Graham DY. Helicobacter pylori and the molecular pathogenesis of intestinal-type gastric carcinoma. Expert Rev Anticancer Ther. 2014;14(8):947–954.
- Danesh J. Helicobacter pylori infection and gastric cancer: systematic review of the epidemiological studies. Aliment Pharmacol Ther. 1999;13(7):851–856.
- Fox JG, Wang TC. Inflammation, atrophy, and gastric cancer. J Clin Invest. 2007;117(1):60–69.
- Hanada K, et al. Helicobacter pylori infection introduces DNA double-strand breaks in host cells. Infect Immun. 2014;82(10):4182–4189.
- Valenzuela MA, Canales J, Corvalán AH, Quest AF. Helicobacter pylori-induced inflammation and epigenetic changes during gastric carcinogenesis. World J Gastroenterol. 2015;21(45):12742–12756.
- Cancer Genome Atlas Research Network . Comprehensive molecular characterization of gastric adenocarcinoma. Nature. 2014;513(7517):202–209.
- Song J, Bent AF. Microbial pathogens trigger host DNA double-strand breaks whose abundance is reduced by plant defense responses. PLoS Pathog. 2014;10(4):e1004030.
- Fedor Y, et al. From single-strand breaks to double-strand breaks during S-phase: a new mode of action of the Escherichia coli Cytolethal Distending Toxin. Cell Microbiol. 2013;15(1):1–15.
- Lord CJ, Ashworth A. The DNA damage response and cancer therapy. Nature. 2012;481(7381):287–294.
- Jeggo PA, Löbrich M. How cancer cells hijack DNA double-strand break repair pathways to gain genomic instability. Biochem J. 2015;471(1):1–11.
- Symington LS, Gautier J. Double-strand break end resection and repair pathway choice. Annu Rev Genet. 2011;45:247–271.
- Lieber MR. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem. 2010;79:181–211.
- Toller IM, et al. Carcinogenic bacterial pathogen Helicobacter pylori triggers DNA double-strand breaks and a DNA damage response in its host cells. Proc Natl Acad Sci U S A. 2011;108(36):14944–14949.
- Chaturvedi R, et al. Activation of EGFR and ERBB2 by Helicobacter pylori results in survival of gastric epithelial cells with DNA damage. Gastroenterology. 2014;146(7):1739–1751.e14.
- Koeppel M, Garcia-Alcalde F, Glowinski F, Schlaermann P, Meyer TF. Helicobacter pylori infection causes characteristic DNA damage patterns in human cells. Cell Rep. 2015;11(11):1703–1713.
- Dorer MS, Fero J, Salama NR. DNA damage triggers genetic exchange in Helicobacter pylori. PLoS Pathog. 2010;6(7):e1001026.
- Sierra JC, et al. Epidermal growth factor receptor inhibition downregulates Helicobacter pylori-induced epithelial inflammatory responses, DNA damage and gastric carcinogenesis. Gut. 2018;67(7):1247–1260.
- Marie MA, Altahir IE. Relationship between the extent of DNA damage and gastritis in normal and Helicobacter pylori-infected patients. Gut Liver. 2011;5(3):315–320.
- Batista PJ, Chang HY. Long noncoding RNAs: cellular address codes in development and disease. Cell. 2013;152(6):1298–1307.
- Schmitt AM, Chang HY. Long noncoding RNAs in cancer pathways. Cancer Cell. 2016;29(4):452–463.
- Ladeira MS, Rodrigues MA, Freire-Maia DV, Salvadori DM. Use of Comet assay to assess DNA damage in patients infected by Helicobacter pylori: comparisons between visual and image analyses. Mutat Res. 2005;586(1):76–86.View this article via: PubMed Google Scholar
- Jeggo PA, Pearl LH, Carr AM. DNA repair, genome stability and cancer: a historical perspective. Nat Rev Cancer. 2016;16(1):35–42.
- Janssen A, Medema RH. Genetic instability: tipping the balance. Oncogene. 2013;32(38):4459–4470.
- Yong X, et al. Helicobacter pylori virulence factor CagA promotes tumorigenesis of gastric cancer via multiple signaling pathways. Cell Commun Signal. 2015;13:30. View this article via: PubMed Google Scholar
- Murata-Kamiya N, et al. Helicobacter pylori CagA interacts with E-cadherin and deregulates the beta-catenin signal that promotes intestinal transdifferentiation in gastric epithelial cells. Oncogene. 2007;26(32):4617–4626.
- Li N, et al. Helicobacter pylori CagA protein negatively regulates autophagy and promotes inflammatory response via c-Met-PI3K/Akt-mTOR signaling pathway. Front Cell Infect Microbiol. 2017;7:417. View this article via: PubMed Google Scholar
- Pierce AJ, Johnson RD, Thompson LH, Jasin M. XRCC3 promotes homology-directed repair of DNA damage in mammalian cells. Genes Dev. 1999;13(20):2633–2638.
- Bennardo N, Cheng A, Huang N, Stark JM. Alternative-NHEJ is a mechanistically distinct pathway of mammalian chromosome break repair. PLoS Genet. 2008;4(6):e1000110.
- Krietsch J, et al. PARP activation regulates the RNA-binding protein NONO in the DNA damage response to DNA double-strand breaks. Nucleic Acids Res. 2012;40(20):10287–10301.
- Li S, et al. Involvement of p54(nrb), a PSF partner protein, in DNA double-strand break repair and radioresistance. Nucleic Acids Res. 2009;37(20):6746–6753.
- Rodrigue A, et al. Interplay between human DNA repair proteins at a unique double-strand break in vivo. EMBO J. 2006;25(1):222–231.
- Salton M, Lerenthal Y, Wang SY, Chen DJ, Shiloh Y. Involvement of Matrin 3 and SFPQ/NONO in the DNA damage response. Cell Cycle. 2010;9(8):1568–1576.
- Ahmed KM, Pandita RK, Singh DK, Hunt CR, Pandita TK. β1-Integrin impacts Rad51 stability and DNA double-strand break repair by homologous recombination. Mol Cell Biol. 2018;38(9):e00672-17. View this article via: PubMed Google Scholar
- Obe G, et al. Chromosomal aberrations: formation, identification and distribution. Mutat Res. 2002;504(1–2):17–36.
- Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–674.
- Grade M, Difilippantonio MJ, Camps J. Patterns of chromosomal aberrations in solid tumors. Recent Results Cancer Res. 2015;200:115–142.
- Yao C, et al. lncRNA TNXA-PS1 modulates Schwann cells by functioning as a competing endogenous RNA following nerve injury. J Neurosci. 2018;38(29):6574–6585.
- Guo CJ, et al. RNA sequencing and bioinformatics analysis implicate the regulatory role of a long noncoding RNA-mRNA network in hepatic stellate cell activation. Cell Physiol Biochem. 2017;42(5):2030–2042.
- Hu X, et al. Profiles of long non-coding RNAs and mRNA expression in human macrophages regulated by interleukin-27. Int J Mol Sci. 2019;20(24):E6207. View this article via: PubMed Google Scholar
- Zhao L, et al. Long non-coding RNA SNHG5 suppresses gastric cancer progression by trapping MTA2 in the cytosol. Oncogene. 2016;35(44):5770–5780.
- Williams GT, Farzaneh F. Are snoRNAs and snoRNA host genes new players in cancer? Nat Rev Cancer. 2012;12(2):84–88.
- Palazzo AF, Lee ES. Non-coding RNA: what is functional and what is junk? Front Genet. 2015;6:2. View this article via: PubMed Google Scholar
- Machado AM, et al. Helicobacter pylori infection induces genetic instability of nuclear and mitochondrial DNA in gastric cells. Clin Cancer Res. 2009;15(9):2995–3002.
- Umeda M, Murata-Kamiya N, Saito Y, Ohba Y, Takahashi M, Hatakeyama M. Helicobacter pylori CagA causes mitotic impairment and induces chromosomal instability. J Biol Chem. 2009;284(33):22166–22172.
- Santos JC, Ribeiro ML. Epigenetic regulation of DNA repair machinery in Helicobacter pylori-induced gastric carcinogenesis. World J Gastroenterol. 2015;21(30):9021–9037.
- Hartung ML, et al. H. pylori-induced DNA strand breaks are introduced by nucleotide excision repair endonucleases and promote NF-κB target gene expression. Cell Rep. 2015;13(1):70–79.
- Lieberman HB. DNA damage repair and response proteins as targets for cancer therapy. Curr Med Chem. 2008;15(4):360–367.
- Powell SN, Bindra RS. Targeting the DNA damage response for cancer therapy. DNA Repair (Amst). 2009;8(9):1153–1165.
- Aparicio T, Baer R, Gautier J. DNA double-strand break repair pathway choice and cancer. DNA Repair (Amst). 2014;19:169–175.
- Srivastava M, Raghavan SC. DNA double-strand break repair inhibitors as cancer therapeutics. Chem Biol. 2015;22(1):17–29.
- Hastings PJ, Lupski JR, Rosenberg SM, Ira G. Mechanisms of change in gene copy number. Nat Rev Genet. 2009;10(8):551–564.
- Hsiao MC, et al. Decoding NF1 intragenic copy-number variations. Am J Hum Genet. 2015;97(2):238–249.
- Lin YL, Gokcumen O. Fine-scale characterization of genomic structural variation in the human genome reveals adaptive and biomedically relevant hotspots. Genome Biol Evol. 2019;11(4):1136–1151.
- Parks MM, Lawrence CE, Raphael BJ. Detecting non-allelic homologous recombination from high-throughput sequencing data. Genome Biol. 2015;16:72. View this article via: PubMed Google Scholar
- Shaw CJ, Lupski JR. Implications of human genome architecture for rearrangement-based disorders: the genomic basis of disease. Hum Mol Genet. 2004;13(spec no 1):R57–R64.View this article via: PubMed Google Scholar
- Kidd JM, et al. A human genome structural variation sequencing resource reveals insights into mutational mechanisms. Cell. 2010;143(5):837–847.
- Bunting SF, Nussenzweig A. End-joining, translocations and cancer. Nat Rev Cancer. 2013;13(7):443–454.
- Smith C, et al. Efficient and allele-specific genome editing of disease loci in human iPSCs. Mol Ther. 2015;23(3):570–577.
- Bibi F, et al. Detection of genetic alterations in gastric cancer patients from Saudi Arabia using comparative genomic hybridization (CGH). PLoS One. 2018;13(9):e0202576.
- Sakugawa ST, et al. API2-MALT1 fusion gene in colorectal lymphoma. Mod Pathol. 2003;16(12):1232–1241.
- Machado AM, et al. Helicobacter pylori infection affects mitochondrial function and DNA repair, thus, mediating genetic instability in gastric cells. Mech Ageing Dev. 2013;134(10):460–466.
- Ajani JA, et al. Gastric Cancer, Version 3.2016, NCCN Clinical Practice Guidelines in Oncology. J Natl Compr Canc Netw. 2016;14(10):1286–1312.View this article via: PubMed Google Scholar
-
Version history
- Version 1 (June 15, 2020): Electronic publication
- Version 2 (June 17, 2020): Corrected figures 2D, 3D, and 3E.
- Version 3 (July 1, 2020): Print issue publication



Copyright © 2024 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)













