Category:Endoplasmic reticulum
Направо към навигацията
Направо към търсенето
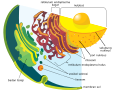
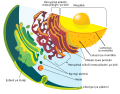
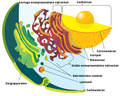
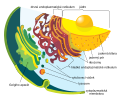
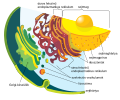
irregular network of membranes coterminous with the outer nuclear membrane in eukaryote cytoplasm that form a meshwork of tubular channels, often expanded into cisternae | |||||
| Качване на файл | |||||
| Екземпляр на |
| ||||
|---|---|---|---|---|---|
| Подклас на |
| ||||
| Част от | |||||
| Връзка с |
| ||||
| |||||
Подкатегории
Показани са 4 от общо 4 подкатегории на тази категория.
E
- Endoplasmic reticulum-associated degradation (0 К, 0 С, 3 Ф)
G
S
- Sarcoplasmic reticulum (0 К, 0 С, 13 Ф)
V
Файлове в категория „Endoplasmic reticulum“
Показани са 200 от общо 202 файла в тази категория.
(предишна страница) (следваща страница)-
0313 Endoplasmic Reticulum a en.png 604 × 362; 230 КБ
-
0313 Endoplasmic Reticulum b en.png 467 × 363; 190 КБ
-
0313 Endoplasmic Reticulum b labeled.png 371 × 363; 196 КБ
-
0313 Endoplasmic Reticulum c en.png 733 × 398; 245 КБ
-
0313 Endoplasmic Reticulum c labeled.png 604 × 398; 250 КБ
-
0313 Endoplasmic Reticulum.jpg 1102 × 892; 488 КБ
-
201601 Endoplasmic reticulum.png 400 × 400; 92 КБ
-
Blausen 0350 EndoplasmicReticulum ar.png 1440 × 1054; 1,21 МБ
-
Blausen 0350 EndoplasmicReticulum.png 1600 × 1054; 1,24 МБ
-
Cellorganeller pic swe 27-05-2007.png 603 × 481; 100 КБ
-
Clara cell lung - TEM.jpg 640 × 480; 98 КБ
-
Drawing of Endoplasmic Reticulum.jpg 2435 × 1807; 1,19 МБ
-
Endomembrane system diagram cs.svg 638 × 530; 326 КБ
-
Endomembrane system diagram de.svg 612 × 486; 96 КБ
-
Endomembrane system diagram en.svg 612 × 486; 109 КБ
-
Endomembrane system diagram es.svg 662 × 506; 95 КБ
-
Endomembrane system diagram fr.svg 612 × 486; 244 КБ
-
Endomembrane system diagram hu.svg 638 × 530; 326 КБ
-
Endomembrane system diagram id.svg 640 × 500; 188 КБ
-
Endomembrane system diagram it.svg 612 × 486; 79 КБ
-
Endomembrane system diagram ka.svg 612 × 486; 89 КБ
-
Endomembrane system diagram ku .svg 612 × 486; 88 КБ
-
Endomembrane system diagram ln.svg 640 × 500; 188 КБ
-
Endomembrane system diagram nl.svg 765 × 600; 192 КБ
-
Endomembrane system diagram notext.svg 560 × 460; 98 КБ
-
Endomembrane system diagram pl.svg 612 × 486; 249 КБ
-
Endomembrane system diagram pt.svg 612 × 486; 286 КБ
-
Endomembrane system diagram ru.svg 612 × 486; 78 КБ
-
Endomembrane system diagram tr.svg 612 × 486; 163 КБ
-
Endomembrane system diagram uk.svg 614 × 498; 295 КБ
-
Endomembrane system diagram zh-tw.svg 612 × 486; 80 КБ
-
Endomembrane system diagram zh.svg 612 × 486; 87 КБ
-
Endoplasmic reticulum - ribosomes.jpg 1221 × 788; 356 КБ
-
Endoplasmic reticulum -- Smart-Servier.jpg 10 240 × 5760; 2,51 МБ
-
Endoplasmic reticulum 1 -- Smart-Servier.png 1755 × 835; 284 КБ
-
Endoplasmic reticulum 2 -- Smart-Servier.png 1898 × 723; 329 КБ
-
Endoplasmic reticulum 3 -- Smart-Servier.png 1900 × 666; 282 КБ
-
Endoplasmic reticulum 4 -- Smart-Servier.png 1760 × 843; 334 КБ
-
Endoplasmic reticulum 5 -- Smart-Servier.png 1749 × 459; 126 КБ
-
Endoplasmic reticulum 6 -- Smart-Servier.png 2117 × 674; 223 КБ
-
Endoplasmic reticulum 7 -- Smart-Servier.png 2118 × 611; 170 КБ
-
Endoplasmic reticulum 8 -- Smart-Servier.png 1753 × 490; 161 КБ
-
-
-
ER-network-dynamics-are-differentially-controlled-by-myosins-XI-K-XI-C-XI-E-XI-I-XI-1-and-XI-2-Movie1.ogv 4,3 сек, 1280 × 720; 2,11 МБ
-
ER-network-dynamics-are-differentially-controlled-by-myosins-XI-K-XI-C-XI-E-XI-I-XI-1-and-XI-2-Movie2.ogv 4,3 сек, 1280 × 720; 2,5 МБ
-
ER-network-dynamics-are-differentially-controlled-by-myosins-XI-K-XI-C-XI-E-XI-I-XI-1-and-XI-2-Movie3.ogv 4,3 сек, 1280 × 720; 1,88 МБ
-
ER-network-dynamics-are-differentially-controlled-by-myosins-XI-K-XI-C-XI-E-XI-I-XI-1-and-XI-2-Movie4.ogv 4,5 сек, 1280 × 720; 2,25 МБ
-
ER-network-dynamics-are-differentially-controlled-by-myosins-XI-K-XI-C-XI-E-XI-I-XI-1-and-XI-2-Movie5.ogv 4,3 сек, 1280 × 720; 1,33 МБ
-
ER-network-dynamics-are-differentially-controlled-by-myosins-XI-K-XI-C-XI-E-XI-I-XI-1-and-XI-2-Movie6.ogv 4,3 сек, 1280 × 720; 1,38 МБ
-
ER-network-dynamics-are-differentially-controlled-by-myosins-XI-K-XI-C-XI-E-XI-I-XI-1-and-XI-2-Movie7.ogv 4,3 сек, 1280 × 720; 1,45 МБ
-
ER-network-dynamics-are-differentially-controlled-by-myosins-XI-K-XI-C-XI-E-XI-I-XI-1-and-XI-2-Movie8.ogv 4,3 сек, 1280 × 720; 1,25 МБ
-
ER-network-dynamics-are-differentially-controlled-by-myosins-XI-K-XI-C-XI-E-XI-I-XI-1-and-XI-2-Movie9.ogv 3,3 сек, 1280 × 720; 683 КБ
-
Esquema de la teoria de l'endosimbio siseriada.png 4408 × 6476; 2,96 МБ
-
Exogenous-Ether-Lipids-Predominantly-Target-Mitochondria-pone.0031342.s006.ogv 28 сек, 480 × 360; 8,69 МБ
-
Golgi-localisation-of-GMAP210-requires-two-distinct-cis-membrane-binding-mechanisms-1741-7007-7-56-S1.ogv 9,0 сек, 453 × 212; 261 КБ
-
-
-
-
-
-
-
-
-
-
-
-
Induction-of-protein-body-formation-in-plant-leaves-by-elastin-like-polypeptide-fusions-1741-7007-7-48-S1.ogv 5,0 сек, 1436 × 1099; 780 КБ
-
Induction-of-protein-body-formation-in-plant-leaves-by-elastin-like-polypeptide-fusions-1741-7007-7-48-S10.ogv 15 сек, 1106 × 1099; 3,34 МБ
-
Induction-of-protein-body-formation-in-plant-leaves-by-elastin-like-polypeptide-fusions-1741-7007-7-48-S11.ogv 3,5 сек, 1106 × 1099; 434 КБ
-
Induction-of-protein-body-formation-in-plant-leaves-by-elastin-like-polypeptide-fusions-1741-7007-7-48-S2.ogv 7,3 сек, 1013 × 1099; 948 КБ
-
Induction-of-protein-body-formation-in-plant-leaves-by-elastin-like-polypeptide-fusions-1741-7007-7-48-S3.ogv 4,7 сек, 1253 × 1099; 792 КБ
-
Induction-of-protein-body-formation-in-plant-leaves-by-elastin-like-polypeptide-fusions-1741-7007-7-48-S4.ogv 6,0 сек, 1106 × 1099; 2,77 МБ
-
-
Induction-of-protein-body-formation-in-plant-leaves-by-elastin-like-polypeptide-fusions-1741-7007-7-48-S6.ogv 13 сек, 1106 × 1099; 2,62 МБ
-
Induction-of-protein-body-formation-in-plant-leaves-by-elastin-like-polypeptide-fusions-1741-7007-7-48-S7.ogv 5,0 сек, 1106 × 1099; 3,55 МБ
-
Induction-of-protein-body-formation-in-plant-leaves-by-elastin-like-polypeptide-fusions-1741-7007-7-48-S8.ogv 5,0 сек, 1106 × 1099; 1,36 МБ
-
Induction-of-protein-body-formation-in-plant-leaves-by-elastin-like-polypeptide-fusions-1741-7007-7-48-S9.ogv 11 сек, 1106 × 1099; 4,15 МБ
-
Inhibition-of-the-Unfolded-Protein-Response-Mechanism-Prevents-Cardiac-Fibrosis-pone.0159682.s006.ogv 1 мин 14 сек, 640 × 368; 39,48 МБ
-
Inside-out-Ca2+-signalling-prompted-by-STIM1-conformational-switch-ncomms8826-s2.ogv 5,3 сек, 1160 × 448; 559 КБ
-
Inside-out-Ca2+-signalling-prompted-by-STIM1-conformational-switch-ncomms8826-s3.ogv 5,0 сек, 752 × 564; 208 КБ
-
Inside-out-Ca2+-signalling-prompted-by-STIM1-conformational-switch-ncomms8826-s4.ogv 5,0 сек, 1004 × 504; 288 КБ
-
Inside-out-Ca2+-signalling-prompted-by-STIM1-conformational-switch-ncomms8826-s5.ogv 6,0 сек, 1176 × 592; 89 КБ
-
-
-
-
-
-
-
-
Mitochondrial-pleomorphy-in-plant-cells-is-driven-by-contiguous-ER-dynamics-Video1.ogv 20 сек, 290 × 184; 1,39 МБ
-
Mitochondrial-pleomorphy-in-plant-cells-is-driven-by-contiguous-ER-dynamics-Video2.ogv 3,8 сек, 548 × 192; 618 КБ
-
Mitochondrial-pleomorphy-in-plant-cells-is-driven-by-contiguous-ER-dynamics-Video3.ogv 7,3 сек, 368 × 258; 970 КБ
-
Mitochondrial-pleomorphy-in-plant-cells-is-driven-by-contiguous-ER-dynamics-Video4.ogv 21 сек, 392 × 602; 1,98 МБ
-
Mitochondrial-pleomorphy-in-plant-cells-is-driven-by-contiguous-ER-dynamics-Video5.ogv 48 сек, 386 × 246; 4,3 МБ
-
Mitochondrial-pleomorphy-in-plant-cells-is-driven-by-contiguous-ER-dynamics-Video6.ogv 45 сек, 378 × 146; 3,25 МБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s016.ogv 6,2 сек, 270 × 235; 145 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s017.ogv 10 сек, 476 × 347; 584 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s018.ogv 3,1 сек, 291 × 230; 141 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s019.ogv 6,7 сек, 216 × 234; 391 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s020.ogv 5,0 сек, 82 × 118; 83 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s021.ogv 9,6 сек, 240 × 283; 166 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s022.ogv 10 сек, 73 × 53; 33 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s023.ogv 4,1 сек, 65 × 89; 26 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s024.ogv 9,3 сек, 58 × 58; 42 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s025.ogv 10 сек, 70 × 86; 64 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s026.ogv 4,0 сек, 68 × 74; 21 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s027.ogv 2,5 сек, 11 × 52; 10 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s028.ogv 36 сек, 68 × 36; 15 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s029.ogv 1,4 сек, 21 × 49; 5 КБ
-
Molecular-Determinants-and-Dynamics-of-Hepatitis-C-Virus-Secretion-ppat.1002466.s030.ogv 5,4 сек, 38 × 50; 19 КБ
-
Motion-and-Flexibility-in-Human-Cytochrome-P450-Aromatase-pone.0032565.s011.ogv 56 сек, 480 × 360; 3,85 МБ
-
Motion-and-Flexibility-in-Human-Cytochrome-P450-Aromatase-pone.0032565.s012.ogv 1 мин 5 сек, 480 × 360; 4,04 МБ
-
Motion-and-Flexibility-in-Human-Cytochrome-P450-Aromatase-pone.0032565.s013.ogv 52 сек, 480 × 360; 6,07 МБ
-
Motion-and-Flexibility-in-Human-Cytochrome-P450-Aromatase-pone.0032565.s014.ogv 1 мин 5 сек, 640 × 360; 15,66 МБ
-
Motion-and-Flexibility-in-Human-Cytochrome-P450-Aromatase-pone.0032565.s015.ogv 1 мин 36 сек, 480 × 360; 8,03 МБ
-
Motion-and-Flexibility-in-Human-Cytochrome-P450-Aromatase-pone.0032565.s016.ogv 1 мин 38 сек, 480 × 360; 6,03 МБ
-
Motion-and-Flexibility-in-Human-Cytochrome-P450-Aromatase-pone.0032565.s017.ogv 2 мин 51 сек, 480 × 360; 14,38 МБ
-
N-linked protein glycosylation in the ER.svg 2662 × 1018; 164 КБ
-
Nuclear-envelope-associated-endosomes-deliver-surface-proteins-to-the-nucleus-ncomms9218-s2.ogv 26 сек, 768 × 576; 4,38 МБ
-
Nuclear-envelope-associated-endosomes-deliver-surface-proteins-to-the-nucleus-ncomms9218-s3.ogv 36 сек, 768 × 576; 2,56 МБ
-
Nuclear-envelope-associated-endosomes-deliver-surface-proteins-to-the-nucleus-ncomms9218-s4.ogv 19 сек, 640 × 480; 6,58 МБ
-
Nuclear-envelope-associated-endosomes-deliver-surface-proteins-to-the-nucleus-ncomms9218-s5.ogv 13 сек, 768 × 576; 4,99 МБ
-
Nucleus ER golgi ex.jpg 534 × 426; 49 КБ
-
Nucleus ER golgi.svg 492 × 565; 168 КБ
-
Nucleus ER.png 416 × 469; 31 КБ
-
Pancreatic acinar cells - TEM.jpg 640 × 480; 160 КБ
-
Pathologies liées aux modifications du réticulum sarcoplasmique.jpg 704 × 449; 120 КБ
-
Phosphatidylinositol hydrolysis and synthesis.jpg 6000 × 4200; 2,65 МБ
-
PLC.png 621 × 706; 276 КБ
-
ProteinTranscription+Synthesis.svg 957 × 531; 82 КБ
-
Rab6-and-Rab11-Regulate-Chlamydia-trachomatis-Development-and-Golgin-84-Dependent-Golgi-ppat.1000615.s006.ogv 5,1 сек, 512 × 512; 1,86 МБ
-
-
Rab6-and-Rab11-Regulate-Chlamydia-trachomatis-Development-and-Golgin-84-Dependent-Golgi-ppat.1000615.s008.ogv 4,2 сек, 512 × 512; 1,38 МБ
-
Rab6-and-Rab11-Regulate-Chlamydia-trachomatis-Development-and-Golgin-84-Dependent-Golgi-ppat.1000615.s009.ogv 5,4 сек, 512 × 512; 2,03 МБ
-
Rab6-and-Rab11-Regulate-Chlamydia-trachomatis-Development-and-Golgin-84-Dependent-Golgi-ppat.1000615.s010.ogv 5,2 сек, 512 × 512; 2,15 МБ
-
-
Rab6-and-Rab11-Regulate-Chlamydia-trachomatis-Development-and-Golgin-84-Dependent-Golgi-ppat.1000615.s012.ogv 5,3 сек, 512 × 512; 1,75 МБ
-
-
-
-
-
-
-
-
-
RER Gland MO.png 1019 × 766; 1,02 МБ
-
Reticle-endoplasmatic.jpg 499 × 291; 53 КБ
-
Reticulo 3D Coanofl.png 786 × 408; 360 КБ
-
Reticulo dinámica Levadu.png 1340 × 1748; 1,03 МБ
-
Reticulo Endoplasmico. Inmunodetección RFP-RPGR..png 856 × 257; 359 КБ
-
Reticulo Levadura.png 357 × 432; 189 КБ
-
Reticulo Nanodominio.png 1200 × 343; 273 КБ
-
Reticulo periferico.png 514 × 754; 378 КБ
-
Reticulo sectores Levad.png 835 × 1341; 951 КБ
-
Retikulum stanica.png 500 × 540; 97 КБ
-
Retículo central Levadura.png 358 × 435; 229 КБ
-
RETÍCULO ENDOPLASMÁTICO.jpg 180 × 239; 37 КБ
-
Retîkûlûma Endoplazmî.png 762 × 418; 221 КБ
-
Rough endoplasmic reticulum.JPG 471 × 278; 33 КБ
-
Rough ER Close up.png 1188 × 672; 138 КБ
-
-
-
-
-
Simultaneous-live-imaging-of-peroxisomes-and-the-ER-in-plant-cells-suggests-contiguity-but-no-Movie6.ogv 8,6 сек, 362 × 138; 549 КБ
-
Smooth Endoplasmic Reticulum.jpg 1195 × 549; 194 КБ
-
Spatial-Temporal-Study-of-Rab1b-Dynamics-and-Function-at-the-ER-Golgi-Interface-pone.0160838.s002.ogv 37 сек, 1920 × 1080; 7,13 МБ
-
Spatial-Temporal-Study-of-Rab1b-Dynamics-and-Function-at-the-ER-Golgi-Interface-pone.0160838.s003.ogv 50 сек, 640 × 360; 776 КБ
-
Spatial-Temporal-Study-of-Rab1b-Dynamics-and-Function-at-the-ER-Golgi-Interface-pone.0160838.s004.ogv 50 сек, 1280 × 720; 4,59 МБ
-
-
The-Effect-of-Gap-Junctional-Coupling-on-the-Spatiotemporal-Patterns-of-Ca2+-Signals-and-the-pcbi.1005295.s001.ogv 1 мин 7 сек, 250 × 103; 1,63 МБ
-
The-Effect-of-Gap-Junctional-Coupling-on-the-Spatiotemporal-Patterns-of-Ca2+-Signals-and-the-pcbi.1005295.s002.ogv 1 мин 7 сек, 100 × 100; 2,42 МБ
-
-
-
-
-
-
-
The-Effect-of-Gap-Junctional-Coupling-on-the-Spatiotemporal-Patterns-of-Ca2+-Signals-and-the-pcbi.1005295.s009.ogv 1 мин 7 сек, 100 × 100; 3,37 МБ
-
The-Effect-of-Gap-Junctional-Coupling-on-the-Spatiotemporal-Patterns-of-Ca2+-Signals-and-the-pcbi.1005295.s010.ogv 1 мин 7 сек, 100 × 100; 1,56 МБ
-
-
-
-
-
-
-
-
The-Human-Polyoma-JC-Virus-Agnoprotein-Acts-as-a-Viroporin-ppat.1000801.s008.ogv 9,1 сек, 180 × 110; 99 КБ
-
-
-
-
-
The-Intracellular-Transport-and-Secretion-of-Calumenin-12-in-Living-Cells-pone.0035344.s005.ogv 5,1 сек, 512 × 512; 5,14 МБ
-
The-Intracellular-Transport-and-Secretion-of-Calumenin-12-in-Living-Cells-pone.0035344.s006.ogv 5,1 сек, 512 × 512; 10,09 МБ
-
The-Intracellular-Transport-and-Secretion-of-Calumenin-12-in-Living-Cells-pone.0035344.s007.ogv 5,0 сек, 512 × 512; 6,93 МБ
-
The-Intracellular-Transport-and-Secretion-of-Calumenin-12-in-Living-Cells-pone.0035344.s008.ogv 5,0 сек, 512 × 512; 13,77 МБ
-
The-Intracellular-Transport-and-Secretion-of-Calumenin-12-in-Living-Cells-pone.0035344.s009.ogv 5,0 сек, 512 × 512; 5,2 МБ